Sampling rivers for genes rather than organisms

Effective environmental management depends on a detailed knowledge of the distribution of species. But taxonomists are in short supply, and some species can be difficult to identify, even for experts. Eawag, in collaboration with Canton Zurich, is now pursuing a new approach for species identification, requiring no more than samples of DNA shed into the environment.
If amphipods are detected in a river, are they a threatened species, or organisms indicating good water quality? Or perhaps the first arrivals of an invasive species? Conservation and environmental management call for a detailed knowledge of species. But experts capable of identifying species under the microscope on the basis of morphological characteristics are increasingly rare. Alternative methods for water monitoring would therefore be welcome. Biologists at Eawag are now pursuing a new approach for the detection of species, involving the use of environmental DNA (eDNA). Because organisms continuously release genetic material into the environment in the form of faeces, hair or skin cells, water samples collected from a river or lake contain innumerable fragments of DNA. As long as the relevant genetic code is known, these DNA segments can be assigned to particular species, using the latest molecular biological techniques and global databases.
Cantonal authorities interested
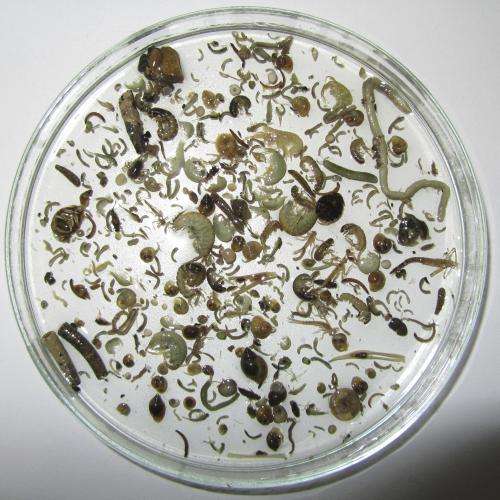
In cooperation with the Canton Zurich Office of Waste, Water, Energy and Air (AWEL), the researchers investigated whether this method is suitable for the detection of macroinvertebrates: organisms such as mayflies, amphipods, mussels or snails are important bioindicators, used in the assessment of water quality and ecotoxicity. Water samples were collected from 14 lake and river habitats in Canton Zurich for eDNA analysis, and macroinvertebrate species collected by kicknet sampling were also determined in the conventional manner.
While the two methods did not always deliver the same results, five of the six target species were reliably detected by both methods. Especially for organisms occurring in small populations, the eDNA method appears to be more sensitive. With this approach, the rare mayfly Baetis buceratus was additionally detected at two sites where no Baetis specimens where found by kicknet sampling. According to project leader Florian Altermatt, the new method may also be suitable for the detection of invasive species at an early stage of colonization. In the US and France, it is already being tested for invasive carp species.
Long-term goal: routine monitoring of biodiversity
The eDNA method offers additional advantages. As eDNA is ubiquitous in freshwater throughout the year, the findings reflect the situation of an entire catchment, and surveillance is less time-critical. By contrast, kicknet sampling merely provides a snapshot, and for many species it can only be carried out at certain stages of the life cycle and certain times of the year. For eDNA analysis, organisms do not have to be removed from a river or lake and – in principle – hundreds of species can be detected at the same time. This means that continuous monitoring of freshwater biodiversity could one day become possible, just as chemical parameters are routinely monitored today.
This, however, is still a long way off: apart from the need for further refinements, the method is still costly and time-consuming. The cantons currently lack the necessary infrastructure and expertise. But Altermatt believes it will not take too long for technical standards to be established, permitting efficient operation: "eDNA analysis will then cost a few hundred Swiss francs and will be cheaper than conventional surveys." However, the new method will not wholly replace the conventional approach. Altermatt argues that the benefits of both approaches should be exploited. In addition, taxonomists will remain indispensable for validation and calibration of the new procedures.
More information: "Utility of environmental DNA for monitoring rare and indicator macroinvertebrate species"; Elvira Mächler, Kristy Deiner, Patrick Steinmann, Florian Altermatt; Freshwater Science, Vol. 33, No. 4 (December 2014), pp. 1174-1183; www.jstor.org/stable/10.1086/678128
Provided by Swiss Federal Institute of Aquatic Science and Technology