Counting fish teeth reveals regulatory DNA changes behind rapid evolution, adaptation

Sticklebacks, the roaches of the fish world, are the ideal animal in which to study the genes that control body shape. They've moved from the ocean into tens of thousands of freshwater streams and lakes around the world, each time changing their skeleton to adapt to the new environment.
Breeding studies between marine and freshwater populations of sticklebacks now have turned up one of the genes that controls tooth number, plus evidence that a simple change in that gene's regulation in a freshwater population is associated with a near doubling in the number of teeth. University of California, Berkeley, scientists say that the corresponding gene in humans may turn out to be involved in tooth, jaw and bone formation.
"This study suggests that the gene, called Bmp6, plays a key role in regeneration of vertebrate organs," said lead researcher Craig Miller, UC Berkeley assistant professor of molecular and cell biology. "Understanding tooth regeneration could lead to a way to replace teeth in humans, for example."
"It's also clear that there is some biological connection between tooth number and cleft palate, because the same regions of the genome control both," he added. "Understanding which genes control the number of teeth is important for understanding what causes malformations, such as a cleft palate."
Miller and his UC Berkeley and Stanford University colleagues reported their findings in last week's issue of the journal Proceedings of the National Academy of Sciences.
The finding has implications, too, for how evolution generates new body shapes. Biologists have proposed that this results primarily from changes in the regulation of a functional gene, not mutations in the gene itself. To date, this has proved true for loss of traits in fish: armor plates, pelvic fins and pigment.
"This is one of the first cases where we find that the rules found for traits lost apply as well to traits gained," said Miller.

Rapid adaptation to fresh water
Like salmon, the two-inch long, threespine stickleback (Gasterosteus aculeatus) is anadromous: it lives in oceans, but swims up freshwater streams to breed. Since the end of the last Ice Age 12,000 years ago, many sticklebacks have colonized lakes and creeks, where their bodies quickly adapted to the new environment: they developed more teeth and stronger jaws, presumably to crack open larger prey found in freshwater, and they lost their armor, perhaps because of fewer predators. In one Alaskan Lake, these changes took a mere 10 years.
Miller, along with his postdoctoral advisor, coauthor David Kingsley of Stanford University, and scientists at the Broad Institute at MIT, sequenced the genomes of sticklebacks from 21 populations in 2012. They and established that all sticklebacks seem to have the same genes, but that rapid change in regulatory DNA allowed them to alter expression of their genes in order to adapt quickly to changing environments.
Because the ancestral saltwater populations still can breed with freshwater populations to produce fertile offspring, researchers can crossbreed fish from different populations to track down the genes and regulatory regions responsible for these body changes. In 2007, shortly after the stickleback genome was sequenced, Miller used crossbreeding to track down genes involved in pigmentation in both fish and humans. At UC Berkeley, he maintains 171 fish tanks with sticklebacks from 11 populations around the world, ranging from Japan and Canada to the San Francisco Bay Area, where 58 of 66 streams are populated with threespine sticklebacks.
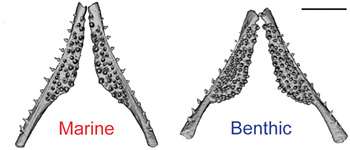
The new study pinpoints one likely gene, bone morphogenetic protein 6 (Bmp6), responsible for tooth number. While the gene seems to be identical in all sticklebacks, regulatory pieces of DNA sitting next to Bmp6 are different in marine versus freshwater fish, suggesting that altered regulation is responsible for the extra teeth. The Bmp6 gene is expressed at higher levels in freshwater fish than in ocean fish.
Goosing up genes
"We think goosing up the Bmp6 signal doubles the number of teeth," Miller said. "This fairly simple genetic basis for gaining something new, such as teeth, is surprising. A very tiny change in the regulator has a large effect."
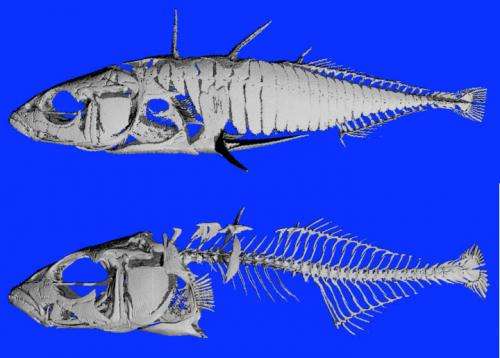
The boost in Bmp6 activity occurs late in the development of the fish larva, when it's already nearly an inch long and halfway to adulthood. Before that time, freshwater and ocean fish have the same number of teeth. The number of teeth in freshwater sticklebacks continues to increase throughout their lifetime as the area of the tooth plate and the density of teeth both increase.
"We found that freshwater-derived sticklebacks keep making teeth constantly and never seem to slow down, whereas the ancestral form stops making more teeth," he said. "While biologists have known for a long time that sharks and some fish continually replace their teeth, almost nothing was known until now about the genetic basis of evolved changes in teeth patterning."
Miller and his colleagues located the Bmp6 gene by crossing marine sticklebacks from Alaska with freshwater sticklebacks from Paxton Lake, Canada. The Canadian fish have about twice the number of teeth as the Alaskan ocean fish. He had earlier identified seven regions of the fish genome involved in controlling tooth number. The new study narrowed down one of these to a stretch of DNA on chromosome 21, which contains the Bmp6 gene.
More information: Evolved tooth gain in sticklebacks is associated with a cis-regulatory allele of Bmp6, Phillip A. Cleves, DOI: 10.1073/pnas.1407567111
Journal information: Proceedings of the National Academy of Sciences
Provided by University of California - Berkeley


















