Biochemist's work helps new company focus on potential hepatitis B cure
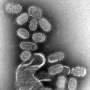
An Indiana University biochemist's discovery of a class of anti-viral small molecules that target the function of a virus DNA hidden in the infected livers of hepatitis B patients may lead to a cure for this viral infection that kills more than 600,000 people annually.
Adam Zlotnick, a professor of molecular and cellular biochemistry in the IU Bloomington College of Arts and Sciences, and four colleagues—chemistry professor Richard DiMarchi and biochemistry visiting scholar William Turner, both of IU Bloomington; Indianapolis biotechnology entrepreneur Derek Small; and infectious disease researcher Dr. Uri Lopatin—formed Assembly Pharmaceuticals in 2012 to develop new anti-viral drugs based on Zlotnick's discoveries. Novel compounds based on these discoveries, known as Core Protein Allosteric Modulators, or CpAMs, are capable of altering the activities of a core hepatitis B protein that is essential for the virus's continued survival.
Despite the early stage of its pipeline, the promise of Assembly's novel approach attracted the interest of Nasdaq-listed Ventrus Biosciences. Last week, Ventrus stockholders voted to merge with Assembly to form a new company, Assembly Biosciences, which is now trading on Nasdaq under the ticker "ASMB," catapulting the firm from new start-up to public company in less than two years.
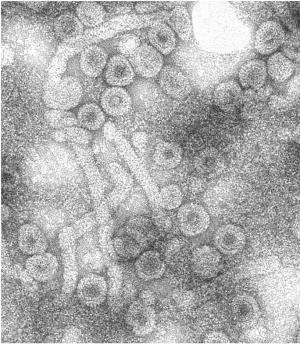
Assembly's anti-viral technology is based on Zlotnick's work at IU focusing on the biophysics of virus self-assembly. Many viruses have a shell, or capsid, that is made of many copies of a virus-specific protein. The Zlotnick lab has shown how purified hepatitis B core protein can spontaneously assemble, in a matter of seconds, to form soccer-ball shaped complexes that are identical to capsids in the infectious virus. By examining the mechanism of assembly, the lab's research has led to the discovery of a number of families of small molecule CpAMs that can selectively and effectively reduce viral load and key viral antigens, considered to be the best marker of a functional cure. Reduction of those viral antigens is and will be a key clinical endpoint in new drug development, Zlotnick said.
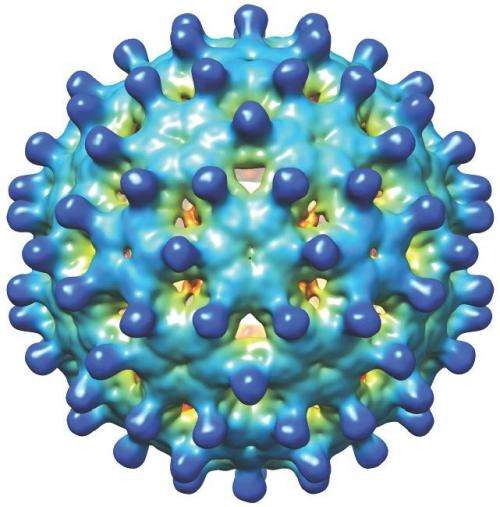
In the case of hepatitis B, CpAMS combat the virus by affecting multiple aspects of the viral lifecycle, including the function of a special viral DNA called covalently closed circular DNA, or cccDNA, that is the viral reservoir hiding in the nuclei of infected liver cells. This cccDNA serves as a template for the production of viral proteins and additional copies of the viral genome, contributing to the persistence of the infection. The cccDNA is not affected by current hepatitis B therapies, which is why only 3 to 5 percent of hepatitis B patients are cured, requiring most infected individuals to take these anti-virals for the rest of their lives.
The hepatitis B virus one of the smallest human pathogens but one of the largest public health problems in the world. As many as 360 million people have chronic hepatitis B, with about 2 million of those in the United States. Chronic infection can lead to cancer and other serious liver diseases, contributing to the estimated 600,000 deaths associated with hepatitis B annually.
One of the key tools used in Zlotnick's work has been IU Bloomington's $1.7 million transmission electron microscope. Supported by funding from the National Science Foundation and the IU College of Arts and Sciences, the microscope allows researchers like Zlotnick to study samples at cryogenic temperatures, where biological specimens are held near the temperature of liquid nitrogen (about negative 196 degrees Celcius), allowing researchers to visualize hydrated samples and determine their structures to near atomic resolution. The Zlotnick lab has used cryo-electron microscopy to examine the protein-protein interactions and the protein-RNA interactions that drive hepatitis B capsid assembly.
Provided by Indiana University