Reconstructing the tree of life

In 1837, Charles Darwin drew a tree of life, a primitive sketch suggesting that all organisms shared a common ancestry. Today, scientists still are trying to reconstruct these evolutionary branches, but using tools, such as genomic data and sophisticated statistical algorithms, that Darwin never could have envisioned.
"We are historians of biology, trying to infer events that happened billions of years ago," says Antonis Rokas, an associate professor of biological sciences at Vanderbilt University. "We take data from what we know today—the DNA of different organisms—and by comparing the sequences and evaluating how similar they are and how different they are from each other, we try to infer the evolutionary relationships between them."
Yet in doing so, phylogeneticists, as they are known, sometimes produce results that are surprising, and even contradictory.
The National Science Foundation (NSF)-funded scientist says it is not unusual for high quality research to report genealogies that conflict with each other over the origins of certain organisms.
"Some are surprising and unexpected, and difficult to decipher," he says.
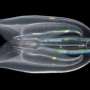
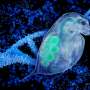
In an attempt to sort out the reasons for the conflicts, and refine the techniques, Rokas and graduate student Leonidas Salichos assembled and analyzed more than 1,000 genes from each of 23 species of yeast, including Saccharomyces cerevisiae, better known as baker's yeast used to make bread, wine and beer, and Candida albicans, sometimes the source of infections.
"Yeasts are a great model for studying ancient branches, since they have very compact genomes, two orders of magnitude smaller than the human genome," Rokas says. "Humans and chimpanzees branched away from each other relatively recently—only 5 or 6 million years ago. With yeast, we're looking at branching events that took place hundreds of millions of years ago."
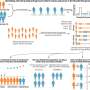
Their study, published in the journal Nature, found that the histories of the more than 1,000 genes all were slightly different from one another, as well as different from the genealogy the researchers built from a simultaneous analysis of all the genes.
"We found 1,070 genes, and made 1,070 trees, and each one was different," Rokas says.
"One explanation may have to do with the fact that you are looking at such a small part of the genome," he adds. "It's like trying to sample the skin color of the United States by looking at only one city. You will get different results if you look at New York, or Nashville, or Washington, D.C."
Rokas and Salichos found that genetic data is less reliable during periods of rapid "radiation," or diversification, when there is a sudden appearance of many new species. "A lot of the debate on the differences in the trees has been between studies concerning the 'bushy' branches that took place in these 'radiations'," Rokas says.
"When you see a lot of conflict, and have lots of data, you expect to see gene differences when you have radiations," he adds. "We don't know what happened. We have 23 yeasts, and what we observed is their DNA sequences in the present. We do these comparisons to try to understand how they came about, and who is most closely related to whom. By looking at these genes, we see consensus in many parts and conflict at the base of the tree. What we understand about evolution leads us to believe that in a small window of time, several new species originated."
The work is important not only because it tackles the enduring mysteries associated with evolution, but, on a practical level, "the process is exactly the same as what we do when we are trying to identify where a new pathogen is coming from," Rokas says. "If there is a new agent of disease or infection, we try to culture it first to see what family of bacteria or viruses it is related to. This allows public health officials to understand very quickly what they are dealing with."
Moreover, "it also allows us to understand the evolution of life on Earth and how a variety of different traits that we associate with different organisms have come about, for example, such characteristics as big brains in humans, compared to other organisms, walking on two legs, loss of hairiness."
Rokas is conducting his research under an NSF Faculty Early Career Development (CAREER) award, which he received in 2009 as part of NSF's American Recovery and Reinvestment Act. The award supports junior faculty who exemplify the role of teacher-scholars through outstanding research, excellent education, and the integration of education and research within the context of the mission of their organization. NSF is funding his work with $688,000 over five years.
The grant's educational goal "is to promote understanding of phylogenetics and its importance for comprehending evolution across the educational continuum," he says, adding that the program has trained three postdoctoral scholars, all of whom have obtained faculty positions, three graduate students, and eight undergraduates, three of whom already have first-author publications. These students include one Hispanic and four women.
The educational component also includes a new undergraduate/graduate course on the computational analysis of genomes; and up to seven advanced graduate training national and international workshops annually, plus numerous lectures at national and international meetings, as well at regional high schools.
If anything, Rokas and his collaborators are discovering that reconstructing the tree of life is anything but simple.
"People expect to find a single tree of life," Rokas says. "They expect there to be one tree that explains how each organism is related to all others."
If that were the case, "then you would expect that different studies would not reach different conclusions," he says. "But you have parts of the tree that are that are easy to infer, where there is consensus, and parts that are challenging. The more ancient the relationships, the harder they are to infer."
His work tries to provide some clarification for why this is the case, "that you should expect to see this when you have these events of rapid diversification, which seemed to have happened rapidly together, at the base of the tree," he says, meaning a very long time ago. "And this means that certain branches of the tree of life will be bushy," he says.
More information: "Inferring ancient divergences requires genes with strong phylogenetic signals." Leonidas Salichos, et al. Nature 497, 327–331 (16 May 2013) DOI: 10.1038/nature12130. Received 06 December 2012 Accepted 28 March 2013 Published online 08 May 2013
Journal information: Nature
Provided by National Science Foundation