Study probes how nanoparticles bind to blood proteins at interfaces
Tiny particles only one millionth of a millimetre across called nanoparticles are abundant in the clothes we wear and even the food we eat. New research published in PCCP indicates that nanoparticles are able to change their binding at surfaces to proteins abundant in the blood depending on whether the protein is bound to fat molecules at the time. The findings indicate how nanoparticles interact with blood proteins in the body by influencing the efficiency of the nanoparticle transport to surfaces.
The work underlies many aspects of protein-nanoparticle adhesion. For example, uncertainty surrounds the safety of nanoparticles in vehicle fumes and a range of everyday products. Toxicologists are concerned that exposure could lead to nanoparticles entering the bloodstream and aggregating in the liver, hindering the functioning of the organ. However, there is also much interest in using nanoparticles in medicine to deliver drugs to specific subcellular regions, such as the nucleus.
In new research, scientists from the Australian National University and the Institut Laue-Langevin (ILL) tested a possible mechanism for nanoparticle binding, known as the 'protein corona' hypothesis. This theory suggests that nanoparticles are able to enter cells because they bind to and become encased by proteins, disguising them from receptors. A key uncertainty was whether this corona structure was also prevalent at surfaces or whether there was different behaviour.
As opposed to many experiments on protein crystals, these experiments were conducted in environments that more closely mimicked human blood. They used silica nanoparticles just 20 nanometers in diameter, similar to those found in industry, in aqueous buffer solutions involving salts at physiological levels to see how they interact with the most abundant protein in our blood, human serum albumin (HSA). The primary role of HSA is to bind to fat molecules in the blood and transport them to different parts of the body, and this binding causes the protein to change shape. Both types of HSA – with and without fat – were studied in this research to investigate if they interacted with the nanoparticles at surfaces differently.
Two complementary experiments were conducted on the buffer-protein-nanoparticle mix to analyse different aspects of the process.
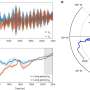
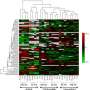
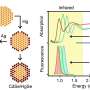
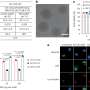
- Neutron reflectometry on the FIGARO instrument at the ILL was used to study how proteins transported the nanoparticles to the air/water interface. Intense beams of neutrons were fired at the surface of the films and the dependence on the angle and the wavelength of the reflected beams provided information about the structure and composition of the different molecules at the interface, and in particular the protein:nanoparticle ratio in the film.
- X-ray reflectometry was used to determine the fine structure of the surface layer, and in particular the distribution of protein molecules decorating the silica nanoparticles at the interface.
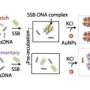
Results showed that several factors are important in the binding. Firstly, the charge on the silica nanoparticle determines how it interacts with protein at surfaces. The silica particles used in the study had a slight negative charge and were attracted to the positively charged domains of HSA even though it also has a net negative charge. Yet the fatted form of the protein has its charge modified by the fat itself, and in that case only the surface interactions were independent of the protein:nanoparticle ratio in the solution. Secondly, the fatted form of the protein is more stable and less likely to unfold. As a result, the protein is less able to transport nanoparticles to the interface in order to adopt optimum conformations at the interface when the effective nanoparticle concentration changes. These results suggest that surface design could be important in minimising toxic effects of nanoparticles and also maximising the therapeutic potential of such particles.
Professor John White, Professor of Physical and Theoretical Chemistry, Research School of Chemistry, Australian National University, says, "As toxic outcomes have been correlated with small size and the problems of particle accumulation the experiments have been done on industrially produced small silica nanoparticles commonly available. They point to stable protein-nanoparticle clustering at interfaces which is sensitive to very subtle properties of the attaching protein. "
Dr Richard Campbell, FIGARO instrument scientist, ILL, says, "A critical part of the research was being able to carry out the measurements on protein molecules in conditions close to their physiological environment. Structural studies on proteins often require the molecule to be in an unnatural crystalline form but the powerful FIGARO reflectometer at the ILL allowed us to study HSA interacting with nanoparticles at the free surface of a buffer solution that more closely mimicked the blood."
Experimental methods
The amount of deuterium – 'heavy hydrogen' – in the buffer solution was altered to exploit a property called isotopic contrast variation. Neutrons are scattered differently by hydrogen and deuterium atoms and by modifying the ratio of H2O to D2O in the buffer the reflection signal from the molecules in question can be enhanced relative to the scattering from the solution. This allows the acquisition of unique structural and composition information that cannot be determined by any other experimental technique.
More information:
"Human Serum Albumin Binding to Silica nanoparticles - effect of protein fatty acid ligand"
John William White, Joo Chuan Ang, Richard A. Campbell, Jack M Lin, Peter N Yaron, Andrew Nelson, Thomas Alured Faunce, Mark Henderson. Phys. Chem. Chem. Phys., 2014, Accepted Manuscript. DOI: 10.1039/C4CP00293H
Provided by Institut Laue-Langevin