Made to order at the synapse: Dynamics of protein synthesis at neuron tip
Protein synthesis in the extensions of nerve cells, called dendrites, underlies long-term memory formation in the brain, among other functions. "Thousands of messenger RNAs reside in dendrites, yet the dynamics of how multiple dendrite messenger RNAs translate into their final proteins remain elusive," says James Eberwine, PhD, professor of Pharmacology, Perelman School of Medicine at the University of Pennsylvania, and co-director of the Penn Genome Frontiers Institute.
Dendrites, which branch from the cell body of the neuron, play a key role in the communication between cells of the nervous system, allowing for many neurons to connect with each other. Dendrites detect the electrical and chemical signals transmitted to the neuron by the axons of other neurons. The synapse is the neuronal structure where this chemical connection is formed, and investigators surmise that it is here where learning and memory occur.
Previous studies in the Eberwine lab have shown that translation of messenger RNAs (mRNAs) into proteins occurs in dendrites at focal points called translational hotspots. Local protein synthesis in dendrites, not in the cell body of nerves, provides the ability to respond rapidly and selectively to external stimuli. This ability is especially important in neurons that have highly polarized cell morphology, meaning one end of the cell has a very different shape from the other end.
In dendrites and axons these rapid structural and functional changes occur concurrently – their length, size, shape, and number change to suit the needs of neuronal cell body communication.
These structural and chemical changes – called synaptic plasticity—require rapid, new synthesis of proteins. Cells may use different rates of translation in different types of mRNA to produce the right amounts and ratios of required proteins.
Knowing how proteins are made to order – as it were - at the synapse can help researchers better understand how memories are made. Nevertheless, the role of this "local" environment in regulating which messenger RNAs are translated into proteins in a neuron's periphery is still a mystery.
Eberwine, first author Tae Kyung Kim, PhD, a postdoc in the Eberwine lab, and colleagues including Jai Yoon Sul, PhD, assistant professor in Pharmacology, showed that protein translation of two dendrite mRNAs is complex in space and time, as reported online in Cell Reports this week.
"We needed to look at more than one RNA at the same time to get a better handle on real- world processes, and this is the first study to do that in a live neuron," Eberwine explains.
At Home in the Hippocampus
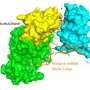
The team looked at two RNAs that make proteins that bind to glutamate, the dominant neurotransmitter in the brain. Using rat hippocampus neurons the researchers found a heterogeneous distribution of translational hotspots along dendrites for the two mRNAs.
This finding indicates that RNA translation is dictated by translational hotspots, not solely when RNA is present. A translational hot spot is characterized by where translation is occurring in a ribosome at any one time in a discrete spot. Since hotspots are not uniform, understanding individual hotspot dynamics is important to understanding learning and memory.
"It's not always one particular RNA that dominates at a translation hotspot versus another type of RNA," says Eberwine. "Since there are 1,000 to 3,000 different mRNA types present in the dendrite overall, but not 1,000 to 3,000 different translational hot spots, do the mRNAs 'take turns' being translated in space and time at the ribosomes at the hotspots?"
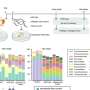
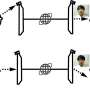
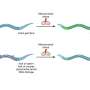
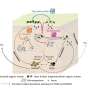
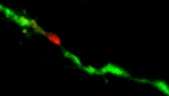
The researchers engineered the glutamate receptor RNAs to contain different fluorescent proteins that are independently detectable, as well as a photo-switchable protein to determine when new proteins were being made. In the case of the photo-switchable protein studies, when an mRNA for the glutamate receptor protein is marked green, it means it has already been translated.
When a laser is passed over the green protein, it changes to red as a way of tagging when it has been been translated, and new proteins synthesized at that hotspot would be green, which is visible by the appearance of yellow fluorescence (green + red, as measured by light on the visible spectrum). These tricks of the light allow the team to keep track of newly made proteins over time and space.
"This is the first time this method of protein labeling has been used to measure the act of translation of multiple proteins over space and time in a quantitative way," says Eberwine. "We call it quantitative functional genomics of live cell translation."
"Our results suggest that the location of the translational hotspot is a regulator of the simultaneous translation of multiple messenger RNAs in nerve cell dendrites and therefore synaptic plasticity," says Sul.
Laying the Groundwork
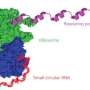
Almost 10 years ago, the Eberwine lab discovered that nerve-cell dendrites have the capacity to splice messenger RNA, a process once believed to take place only in the nucleus of cells. Here, a gene is copied into mRNA, which possesses both exons (mature mRNA regions that code for proteins) and introns (non-coding regions). mRNA splicing works by cutting out introns and merging the remaining exon pieces, resulting in an mRNA capable of being translated into a specific protein.
The vast array of proteins within the human body arises in part from the many ways that mRNAs can be spliced and reconnected. Specifically, splicing removes pieces of intron and exon regions from the RNA. The resulting spliced RNA is made into protein.
If the RNA has different exons spliced in and out of it, then different proteins can be made from this RNA. The Eberwine lab was successful in showing that splicing can occur in dendrites because they used sensitive technologies developed in their lab, which permits them to detect and quantify RNA splicing, as well as the translated protein in single isolated dendrites.
Understanding the dynamics of RNA biology and protein translation in dendrites promises to provide insight into regulatory mechanisms that may be modulated for therapeutic purposes in neurological and psychiatric illnesses. The directed development of therapeutics requires this detailed knowledge, says Eberwine.
Journal information: Cell Reports
Provided by University of Pennsylvania School of Medicine