Researchers develop new sensor for methylated DNA
Collaborators from Mayo-Illinois Alliance for Technology Based Healthcare have developed a new, single molecule test for detecting methylated DNA. Methylation—the addition of a methyl group of molecules to a DNA strand—is one of the ways gene expression is regulated. The findings appear in the current issue of Scientific Reports.
"While nanopores have been studied for genomic sequencing and screening analysis, this new assay can potentially circumvent the need for some of the current processes in evaluating epigenetics-related diseases," says George Vasmatzis, Ph.D., co-leader of Mayo's Biomarker Discovery Program in the Center for Individualized Medicine and co-lead author on the article. He says the assay could eliminate the need for bisulfite conversion of DNA, fluorescent labeling, and polymerase chain reaction (PCR).
"Next steps include increasing the spatial resolution by incorporating thinner membranes and by integrating the same preparation steps," says Rashid Bashir, Ph.D., professor of bioengineering, director of the Micro and Nanotechnology Laboratory, and co-lead author of the study at the University of Illinois at Urbana-Champaign.
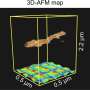
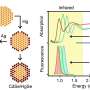
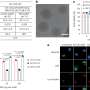
A nanopore, in this case, is a very small hole in an artificial membrane, that allows only a single molecule to be located and identified. Researchers say this is useful as methylation in promoter sequences can indicate tumor development in most major types of cancer and may be a better biomarker than many genetic markers. Scientists are now able to differentiate methylated from non-methylated DNA by attaching a protein on the methylated nucleotides measuring ionic electrical current via a solid-state nanopore.
Journal information: Scientific Reports
Provided by Mayo Clinic