Fruit fly studies guide investigators to misregulated mechanism in human cancers
Changes in how DNA interacts with histones—the proteins that package DNA—regulate many fundamental cell activities from stem cells maturing into a specific body cell type or blood cells becoming leukemic. These interactions are governed by a biochemical tug of war between repressors and activators, which chemically modify histones signaling them to clamp down tighter on DNA or move aside and allow a gene to be expressed.
In the November 19, 2012, online edition of the journal Genes & Development, researchers at the Stowers Institute for Medical Research report findings that may unveil the role of two human genes—MLL3 and MLL4—that are frequently mutated in certain cancers. In addition to its disease implications, the Stowers study exemplifies how the analysis of model organisms like yeast and fruit flies can illuminate human molecular biology.
"We know that fundamental regulatory machineries are highly conserved from yeast to Drosophila to humans," says Stowers Investigator and the study's senior author, Ali Shilatifard, Ph.D., whose lab website describes projects in yeast, fruit flies, and mammalian species. "We use the awesome power of yeast and Drosophila genetics and biochemistry to define the molecular properties of these fundamentally important factors, and then test their function in human cells. In this study, these genes turned out to be frequently mutated in several different types of cancers."
Over a decade ago, studies conducted by Shilatifard's laboratory identified an assembly of proteins known as Set1/COMPASS that biochemically modifies Histone 3 (H3) by planting methyl groups at a very precise location on the histone—namely lysine 4 (K4)—within chromosomes. H3K4 can be mono-, di-, or trimethylated by Set1/COMPASS. H3K4 trimethylation by Set1/COMPASS has now become the hallmark of actively transcribed genes from yeast to human. H3K4 can also be monomethylated, and this modification seems to be specific to enhancers, which are DNA elements that regulate gene expression in a tissue specific manner.
"After we discovered yeast COMPASS, we found that human cells have the same machinery," says Shilatifard. "But rather than one COMPASS methylase, human cells bear six. The question became, why are there six COMPASS family members in human cells?" However, it has not been clear which COMPASS family member functions as monomethylase on enhancers.
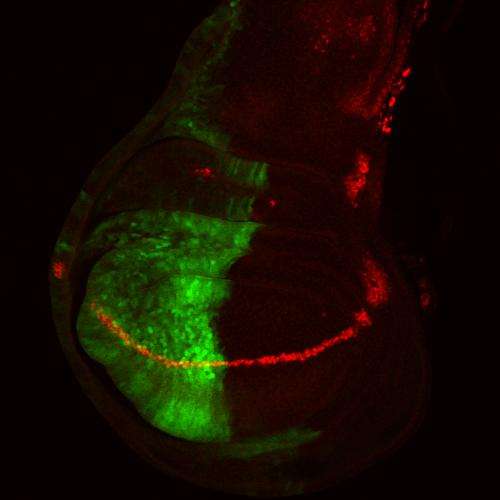
The first clue came from Shilatifard's 2011 Molecular and Cellular Biology study reporting that cells from the fruit fly Drosophila contained three COMPASS family members. In the new study, the group, led by postdoctoral fellow Hans-Martin Herz, Ph.D., unveiled the true fly "monomethylator" by blocking the expression of the components of each of the three candidates and imaging Drosophila tissues to assess methylation of H3K4. Attenuating the fly COMPASS-related complex called Trr (for Trithorax-related) caused genome-wide decreases in H3K4 monomethylation, while disabling the other two candidates did not.
To further the case, Herz collaborated with bioinformatician Alexander Garruss to examine where Trr "sits" genome-wide on fly genes and discovered that it associated with DNA regions called enhancer, the activation zones often flanking a target gene. Enhancers are required to switch genes on, a process Shilatifard likens to how an airline pilot commands his control panel by turning on and off switches to get the plane to move forward and gather enough speed to take off. "We found that Trr needs to move to an enhancer for it to become active," says Herz. "That suggests that Trr's presence orchestrates the transition from inactive to active enhancers."
Two mammalian COMPASS-like complexes structurally resemble Trr, which include proteins called MLL3 and MLL4. Focusing on MLL3, the group cultured mouse embryonic cells lacking MLL3 and undertook a global histone methylation analysis to observe where changes in methylation patterns occurred. They found that H3K4 monomethylation was reduced in enhancer regions.
"MLL3 and MLL4 are mutated in numerous cancers," says Shilatifard. "Researchers have found that many lymphomas have mutations in MLL3 and MLL4."
Herz agrees, noting that mutations in MLL3 and MLL4 are seen in human colorectal cancer, medulloblastoma, breast tumors, and leukemia. "This suggests that activation and deactivation of enhancers could play an important role in cancer pathogenesis," he says. "Understanding which proteins carry out this function or how they control enhancer activity could help us to understand how genes that suppress tumor suppressor genes are deactivated in various cancers."
Journal information: Genes & Development , Molecular and Cellular Biology
Provided by Stowers Institute for Medical Research