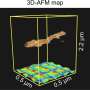
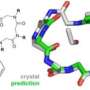
New insights into membrane-assisted self-assembly
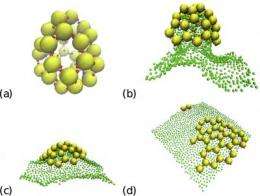
How proteins and virus capsids - complex protein structures that encase the genetic material of viruses - form structures near to a fluctuating membrane is simulated by physicist Richard Matthews with advanced computational techniques. Matthews performs research as a Lise-Meitner-Fellow in the Computational Physics Group of the University of Vienna under the direction of Christos Likos, professor for Multiscale Computational Physics. The results are relevant to the understanding of biophysical processes and appear in the current issue of Physical Review Letters.
"In our current paper we present new computational results that explore how membranes may influence crucial biological processes", explains Richard Matthews, Lise-Meitner-Fellow at the University of Vienna and first author of the study. The focus of the investigation is the self-assembly of microscopic particles, the formation of structures or patterns without human intervention. More specifically, the effect of the interactions between membranes and proteins, which can influence the formation of ordered structures in cells, is considered.
Self-assembly has become a hot topic in recent years. Many of the most astonishing examples are found in nature, from tiny motors (e.g. the flagellum motor) to virus capsids with perfect spherical forms. Many researchers have also tried to improve our understanding by representing the assembly of such structures with models. In order to gain clear insight it is preferable that these models are as simple as possible. This approach has been very successful in reproducing key features of experiments, whilst also uncovering new aspects. In reality, these processes do not occur in isolation and, in fact, many happen on, or in the vicinity of, membranes, a fact that has been previously neglected in the construction of simple models.
Advanced Simulation Techniques
The research aims to find out the general properties of these fascinating systems by applying state of the art simulation techniques. This requires everything to be calculated on a computer. Due to the complexity of the task, high performance computers are necessary. "In our work we have applied advanced simulation techniques, which allowed us to see how interactions with a membrane influence self-assembly", explains Richard Matthews. "We determined that membranes promote self-assembly and also find that our model reproduces structures that are very similar to those seen in nature."
More information: Physical Review Letters. October 2012. DOI: 10.1103/PhysRevLett.109.178302
Journal information: Physical Review Letters
Provided by University of Vienna