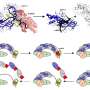
A long-held assumption confirmed: We can learn a lot from other species' genes
Researchers at the SIB Swiss Institute of Bioinformatics and the EMBL-European Bioinformatics Institute have confirmed the long-held belief that studying the genes we share with other animals is useful. The study, published today in the open access journal PLoS Computational Biology, shows how bioinformatics makes it possible to test the fundamental principles on which life science is built.
Studying genes helps life science researchers understand how our bodies work and how diseases progress. Scientists have long looked to model species – mice, for example – to understand human biology. This is at the root of what is called the 'ortholog conjecture': the idea that we can take what we learn from a few species and apply it to many.
The ortholog conjecture
To get an idea of what orthologs are about, consider wolf teeth. If we want to know more about our canine teeth, would we learn more by looking at the canines of wolves? Or would it be better to look at our molars? The answer might not be straightforward. In genetics, scientists address a similar question: Is it better to compare genes in mice and humans that directly descend from a common ancestor (these are called 'orthologs') – or to compare imperfect copies of genes within a human being (the 'paralogs')?
Assume nothing
For the past 40 years, scientists have gone with Plan A: the orthologs, and this has worked quite well. Studying genes in model species has provided invaluable insights in all areas of biology. But until now, there hasn't been enough data to answer this question with authority. With advances in biotechnology producing vast quantities of data every day, there is finally enough to settle the debate.
Using advanced computational techniques on data derived from tens of thousands of scientific articles, the researchers analysed 400 000 pairs of genes (orthologs and paralogs) from 13 different species. The team compared the two approaches and picked a winner.
"We have the data to prove that the study of orthologs is indeed useful, but we are only at the beginning," says Prof. Marc Robinson-Rechavi of SIB and the University of Lausanne. "This is at the heart of all of comparative genomics, in which we try to extrapolate knowledge from a handful of organisms and apply it to all of life."
"We found that current experimental annotations do support the standard model," explains Christophe Dessimoz of EMBL-EBI. "Our work corroborates the assumption that studying the genes of other species – whether mice, yeast, or even bacteria – can elucidate aspects of human biology."
The same question has recently been addressed by Matthew Hahn and colleagues (University of Indiana, USA), whose different conclusion sparked some debate. The new research demonstrates that these controversial results were due to overlooked biases in the collective knowledge of gene function. Controlling for these, the new study unequivocally supports the ortholog conjecture and the fact that studying species we are only distantly related to – even worms, flies, yeasts or bacteria – is relevant and useful.
Open science
This study was made possible by the tradition of open science in bioinformatics, which is strongly supported by SIB, EMBL-EBI and ELIXIR, the incipient infrastructure for life science data in Europe. All of the data used in the study was freely available, including the genome sequences and experimental knowledge described in thousands of publications. ELIXIR will build on this tradition and provide the next generation of infrastructure for biological information in Europe and worldwide.
More information: Adrian M. Altenhoff, Romain A. Studer, Marc Robinson-Rechavi and Christophe Dessimoz. (2012) Resolving the ortholog conjecture: orthologs tend to be weakly, but significantly, more similar in function than paralogs. PLoS Comp Biol (in press). DOI:10.1371/journal.pcbi.1002514
Journal information: PLoS Computational Biology
Provided by Swiss Institute of Bioinformatics