A matter of priorities: Bacteria evolved way to safeguard crucial genetic material

Just as banks store away only the most valuable possessions in the most secure safes, cells prioritise which genes they guard most closely, researchers at the European Molecular Biology Laboratory’s European Bioinformatics Institute (EMBL-EBI) have found. The study, published online today in Nature, shows that bacteria have evolved a mechanism that protects important genes from random mutation, effectively reducing the risk of self-destruction. The findings answer a question that has been under debate for half a century and provide insights into how disease-causing mutations arise and pathogens evolve.
“We discovered that there must be a molecular mechanism that preferentially protects certain areas of the genome over others,” says Nicholas Luscombe, who led the research at EMBL-EBI. “If we can identify the proteins involved and uncover how this works, we will be even closer to understanding how mutations that lead to diseases like cancer can be prevented.”
Mutations are the reason each of us is unique. These changes to our genetic material are at the root of variation between individuals, and between cells within individuals. But they also have a darker side. If it affects an important gene – for example, rendering a tumour-suppressing gene useless – a mutation can have disastrous consequences. Nevertheless, protecting all genes from mutation would use up too many of the cell’s resources, just like holding all deposits in maximum-security safes would be prohibitively expensive. Iñigo Martincorena, a PhD student in Luscombe’s lab, has now found that cells evolved a ‘risk management’ strategy to address this issue.
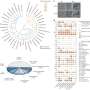
Looking at 120 000 tiny genetic mutations called single nucleotide polymorphisms (SNPs) in 34 strains of the bacterium E. coli, the scientists were able to quantify how random the mutation rate was in different areas of the bacterial genomes. Their results showed that key genes mutate at a much lower rate than the rest of the genetic material, which decreases the risk of such genes suffering a detrimental mutation.
“We were struck by how variable the mutation rate appears to be along the genome,” says Martincorena. “Our observations suggest these bacteria have evolved a clever mechanism to control the rate of evolution in crucial areas of the genome.”
Using population genetics techniques, the researchers were able to disentangle the effects of mutation rate and natural selection on mutations, settling a long-standing debate in the field. Scientists have long thought that the chances of a mutation occurring were independent of its value to an organism. Once the mutation had occurred, it would undergo natural selection, spreading through the population or being eliminated depending on how beneficial or detrimental the genetic change proved to be.
“For many years in evolution there has been an assumption that mutations occur randomly, and that selection ‘cleans them up’,” explains Martincorena. “But what we see here suggests that genomes have developed mechanisms to avoid mutations in regions that are more valuable than others.”
Observations from studies of cancer genomes suggest that similar mechanisms may be involved in the development of cancers, so Luscombe and colleagues would now like to investigate exactly how this risk-managing gene protection works at a molecular level, and what role it may play in tumour cells.
More information: Martincorena, I., Seshasayee, A.S.N., and Luscombe, N. (2012). Evidence of non-random mutation rates suggests an evolutionary risk management strategy. Nature, Advance Online Publication 22 April 2012. DOI: 10.1038/nature10995
Abstract
A central tenet in evolutionary theory is that mutations occur randomly with respect to their value to an organism; selection then governs whether they are fixed in a population. This principle has been challenged by long-standing theoretical models predicting that selection could modulate the rate of mutation itself1, 2. However, our understanding of how the mutation rate varies between different sites within a genome has been hindered by technical difficulties in measuring it. Here we present a study that overcomes previous limitations by combining phylogenetic and population genetic techniques. Upon comparing 34 Escherichia coli genomes, we observe that the neutral mutation rate varies by more than an order of magnitude across 2,659 genes, with mutational hot and cold spots spanning several kilobases. Importantly, the variation is not random: we detect a lower rate in highly expressed genes and in those undergoing stronger purifying selection. Our observations suggest that the mutation rate has been evolutionarily optimized to reduce the risk of deleterious mutations. Current knowledge of factors influencing the mutation rate—including transcription-coupled repair and context-dependent mutagenesis—do not explain these observations, indicating that additional mechanisms must be involved. The findings have important implications for our understanding of evolution and the control of mutations.
Journal information: Nature
Provided by European Molecular Biology Laboratory