HeliScopeCAGE: A new gene expression analysis technique on a single molecule sequencer
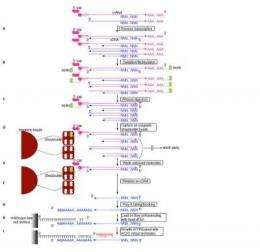
A new gene expression technique adapted for single molecule sequencing has enabled researchers at the RIKEN Omics Science Center (OSC) to accurately and quantitatively measure gene expression levels using only 100 nanograms of total RNA. The technique, which pairs RIKEN's Cap Analysis of Gene Expression (CAGE) protocol with the Helicos® Genetic Analysis System developed by Helicos BioSciences Corporation, opens the door to the detailed analysis of gene expression networks and rare cell populations.
In recent years, next-generation DNA sequencers have produced an increasingly detailed picture of how genes are expressed at the molecular level. The transcriptional output of these genes – the RNA copies produced from DNA – has revealed a richness of complexity in transcript structure and function, providing insights into the molecular-level properties of cancers and other diseases.
One of the most powerful methods for analyzing RNA transcripts is the Cap Analysis of Gene Expression (CAGE) protocol developed at the RIKEN OSC. A unique approach, CAGE enables not only high-throughput gene expression profiling, but also simultaneous identification of transcriptional start sites (TSS) specific to each tissue, cell or condition.
With HeliScopeCAGE, the OSC research team has adapted the existing CAGE protocol for use with the revolutionary HeliScopeTM Single Molecule Sequencer. Unlike earlier sequencers, the HeliScope Sequencer does not employ polymerase chain reaction (PCR) amplification to multiply a small number of DNA strands for analysis, a process which can introduce biases into data. Instead, the HeliScope Sequencer actually sequences the DNA strand itself, enabling direct, high-precision measurement.
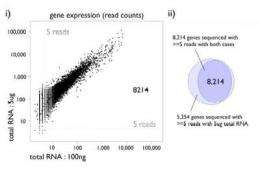
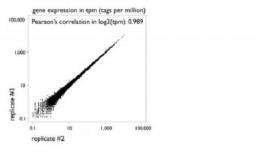
In a paper published in Genome Research, RIKEN researchers confirm that this direct approach reduces biases and generates highly reproducible data from between 5 micrograms to as little as 100 nanograms of total RNA. A comparison using a leukemia cell line (THP-1) and a human cervical cancer cell line (HeLa) further shows that results from the technique are closely correlated to those from traditional microarray analysis. By making possible high-precision gene expression analysis from tiny samples, HeliScopeCAGE greatly expands the scope of research at the OSC, strengthening the institute's role in Japan as a hub for next-generation genome analysis.
Provided by RIKEN