Basic peptides take a different route to the same destination
Flouting expectations is what a small, slightly basic protein does when it comes to breaking up, according to scientists at Pacific Northwest National Laboratory and the University of Hong Kong. Using a specialized mass spectrometer and a technique that involves propelling proteins into a hard surface, the scientists found that a certain small protein or peptide ignores the conventional approach to fragmentation, and takes a completely different path.
In laboratories around the world, new proteins and those indicative of disease in medical tests are identified by snipping the unknown proteins apart using a technique that creates pieces called peptides that often end in the amino acid arginine. Then, the scientists must identify the peptides. It is like shredding a piece of paper, without looking at it, and then putting it back together.
"Unfortunately, we can only identify 20 to 25 percent of the peptides we get. The rest are not assigned, even when we have good quality data from a mass spectrometer," said Dr. Julia Laskin, the principal investigator and a physical chemist at Pacific Northwest National Laboratory. "We want to improve that number by better understanding the rules."
The researchers wanted to see how a small, basic arginine-terminated peptide broke apart. For years, experimental scientists showed the peptide followed a mobile proton or conventional pathway to breaking bonds. But theoreticians suspected another route. The team decided to see if they could prove this other route existed and determine when the commonly seen arginine-based peptide selected this alternative pathway.
The key factors peptides use in selecting a route are energy barriers and entropy effects. Energy barriers are the amount of heat, light, or other source of power that must be added to make something happen. Entropy effects are the amount of chaos or disorder needed for a reaction to happen.
With the peptides created, the team used surface-induced dissociation on a specially configured Fourier transform ion cyclotron resonance mass spectrometer, located in DOE's EMSL. This specialized technique fired peptides at a surface. The peptides hit the surface and bounced back as fragments. Using this technique and sophisticated modeling, the researchers measured the energy and entropy needed to break apart the molecule.
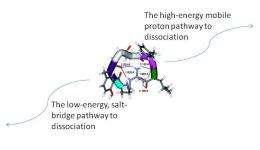
Based on the experimental and modeling results, the team demonstrated that arginine-terminated peptides use the unexpected pathways, which involve structures known as salt bridge intermediates. This pathway doesn't require a lot of energy, but the entropy or disorder decreases and significantly slows down this reaction. The entropy does not favor fragmentation because the molecule must jostle and bump into just the perfect conformation to break apart. In contrast, when the molecule breaks along the conventional mobile proton pathway the energy barrier is high but the favorable entropy factor speeds up the reaction.
So, entropy is critical for fragmentation, as is the amount of energy available.
"The involvement of alternative dissociation channels for arginine-containing peptides was predicted by the theory group, but now we've showed it experimentally," said Laskin. "It was almost impossible to measure this fragmentation using conventional experimental techniques, until we used the surface-induced dissociation approach."
This research is part of a broader effort to understand how peptides fragment. The team is now looking to bring in theory and advanced computational capabilities to complement the experimental research results.
More information: Laskin J, et al. 2010. "Effect of the basic residue on the energetics, dynamics, and mechanisms of gas-phase fragmentation of protonated peptides." Journal of the American Chemical Society 132(45):16006-16016.
Provided by Pacific Northwest National Laboratory


















