Bacteria Are Better Gene Packers Than We Thought
(PhysOrg.com) -- In microbial genomes, genes are typically depicted as linear series of separate regulatory and coding regions. This leads to the assumption that annotations done by computer to predict such arrangements completely describe the coding capacity of bacterial genomes.
However, the more complex organisms such as plants and animals pack their genes into their DNA very densely. One common packing trick is to code genes on both strands of the DNA, allowing the genes to overlap along the chromosome. Bacterial genes previously shown to reside on the second, or anti-sense, strand overlap just a little -- a couple dozen DNA bases, for example. Less than a handful of genes overlap completely.
To test whether bacteria might be packing genes more tightly than this, scientists at Pacific Northwest National Laboratory and Tufts University compared the proteins made by a bacterial species with what is known about its genome. They chose Pseudomonas fluorescens, Gram-negative rod-shaped bacteria that inhabit soil, plants, and water surfaces.
The researchers analyzed these proteins at the U.S. Department of Energy's EMSL, a national scientific user facility at PNNL, using ultra high-pressure reversed-phase high-performance liquid chromatography coupled to an ion trap mass spectrometer.
Using all the information from a 6-frame translation of the bacterial genome, the team identified as many proteins made by P. fluorescens as their instruments would allow and deduced the gene sequence needed to create those proteins. Then they compared the deduced genes to the genes found by the annotation of the genome.
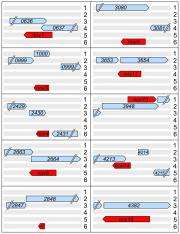
The team subsequently analyzed coding sequences, reading frames, and comparative alignments and found 16 genes for proteins not previously mapped to the P. fluorescens genome. The researchers found nine previously unknown genes coded on the anti-sense strand of DNA. Unlike other anti-sense genes found in bacteria, however, these genes overlapped other ones on the sense strand completely or nearly so. This suggests that researchers have under-estimated how often bacteria pack genes by overlapping them.
The results suggest that the cues researchers use to identify genes by sequence in a stretch of DNA have not all been identified. The 16 newly identified genes improve the quality of the Pf0-1 genome annotation, and the detection of anti-sense protein-coding genes indicates the underappreciated complexity of bacterial genome organization.
The results show that tools currently used to identify the complete set of genes and proteins in organisms, especially bacteria, are insufficient. But work such as this will lead to a more comprehensive understanding of how the genomic blueprint within bacteria translates into functioning proteins that converge into a living organism. Such an understanding could also lead to insights in evolutionary biology.
This work was supported by DOE's Office of Biological and Environmental Research's Genomics Science Program. The research team includes Kim Wook, Mark Silby, Julie Nicoll, and Stuart Levy, Tufts; and Sam Purvine, Kim Hixson, Matt Monroe, Carrie Nicora, and Mary Lipton, PNNL.
More information: Wook K, MW Silby, SO Purvine, JS Nicoll, KK Hixson, ME Monroe, CD Nicora, MS Lipton, and SB Levy. 2009. "Proteomic Detection of Non-Annotated Protein-Coding Genes in Pseudomonas fluorescens Pf0-1," PLoS ONE 4(12):e8455. doi:10.1371/journal.pone.0008455
Provided by Pacific Northwest National Laboratory