Computer simulations reveal the energy landscape of ion channels
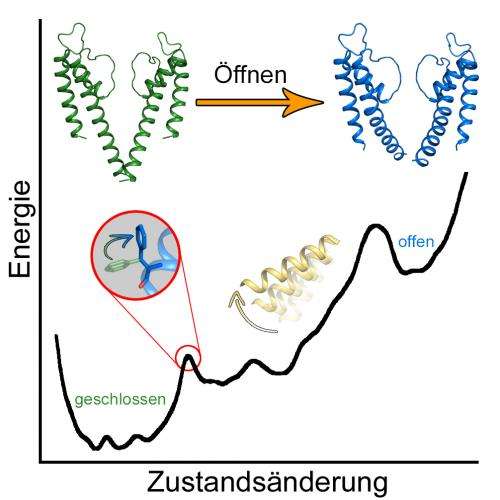
Every cell of our body is separated from its environment by a lipid bilayer. In order to maintain their biological function and to transduce signals, special proteins, so called ion channels, are embedded in the membrane. Anna Stary-Weinzinger and Tobias Linder from the University of Vienna and Bert de Groot from the Max Planck Institute of Biophysical Chemistry in Göttingen identified a key amino acid (phenylalanine 114), which plays an essential role for opening and closing of these ion channels. A conformational change of phenylalanine triggers opening of the channels.
"These proteins are highly selective, they can distinguish between different ions such as sodium, potassium or chloride and allow ion flux rates of up to 100 million ions per seconds", explains Stary-Weinzinger, leader of the research project and postdoc at the Department of Pharmacology and Toxicology of the University of Vienna. "These molecular switches regulate numerous essential body functions such as transduction of nerve signals, regulations of the heart rhythm or release of neurotransmitters. Slight changes in function, caused by replacement of single amino acids, can lead to severe diseases, such as arrhythmias, migraine, diabetes or cancer."
Knowledge of ion channel function provides the basis for better drugs
Ion channels are important drug targets. 10 percent of current pharmaceuticals target ion channels. A detailed understanding of these proteins is therefore essential to develop drugs with improved risk-benefit profiles. An important basis for drug development is a detailed knowledge of the functional mechanisms of these channels. However, there are still many open questions; especially the energy profile and pathway of opening and closure are far from being understood.
Computer simulations visualize ion channel movements
To watch these fascinating proteins at work, molecular dynamics simulations are necessary. Computational extensive calculations were performed with the help of the Vienna Scientific Cluster (VSC), the fastest high performance computer in Austria, a computer cluster operated by the University of Vienna, the Vienna University of Technology and the University of Natural Resources and Applied Life Sciences Vienna. With the help of VSC, the free energy landscape of ion channel gating could be investigated for the first time. The young researchers discovered that the open and closed channel states are separated by two energy barriers of different height.
Phenylalanine triggers conformational changes
Surprisingly, the dynamics of a specific amino acid, phenylalanine 114, are coupled to a first smaller energy barrier. "This side chain acts as molecular switch to release the channel from the closed state", explains Tobias Linder, PhD student from the University of Vienna. After these local changes, the channel undergoes large global rearrangements, leading to a fully open state. This second transition from an intermediate to a fully open pore is accompanied by a large second energy barrier.
Publication details
T. Linder, B.L. de Groot, A. Stary-Weinzinger: probing the energy landscape of activation gating of the bacterial potassium channel KcsA. PLOS Computational Biology, May 2013. DOI: 10.1371/journal.pcbi.1003058
Journal information: PLoS Computational Biology
Provided by University of Vienna