Researchers develop new tool for making genetic engineering of microbial circuits reliably predictable
(Phys.org)—Synthetic biology is the latest and most advanced phase of genetic engineering, holding great promise for helping to solve some of the world's most intractable problems, including the sustainable production of energy fuels and critical medical drugs, and the safe removal of toxic and radioactive waste from the environment. However, for synthetic biology to reach its promise, the design and construction of biological systems must be as predictable as the assembly of computer hardware.
An important step towards attaining a higher degree of predictability in synthetic biology has been taken by a group of researchers with the U.S. Department of Energy's Lawrence Berkeley National Laboratory (Berkeley Lab) under the leadership of computational biologist Adam Arkin. Arkin and his team have developed an "adaptor" that makes the genetic engineering of microbial components substantially easier and more predictable by converting regulators of translation into regulators of transcription in Escherichia coli. Transcription and translation make up the two-step process by which the coded instructions of genes are used to synthesize proteins.
"Application of our adaptor should produce large collections of transcriptional regulators whose inherent composability can facilitate the predictable engineering of complex biological circuits in microorganisms," Arkin says. "This in turn should allow for safer and more efficient constructions of increasingly complex functions in microorganisms."
Arkin is the director of Berkeley Lab's Physical Biosciences Division and the corresponding author of a paper describing this work in Nature Methods. The paper is titled "An adaptor from translational to transcriptional control enables predictable assembly of complex regulation. Co-authoring this paper were Chang Liu, Lei Qi, Julius Lucks, Thomas Segall-Shapiro, Denise Wang and Vivek Mutalik.

Synthetic biology combines modern principles of science and engineering to develop novel biological functions and systems that can tackle problems natural systems cannot. The focus is on bacteria and other microbes that can metabolize a wide variety of valuable chemicals and molecules, and play a critical role in the global cycles of carbon and other important elements. One of the keys to success in synthetic biology is the design and construction of customized genetic switches in microbes that can control the expression of both coding and non-coding RNA, act on operons (small groups of genes with related functions that are co-transcribed in a single strand of messenger RNA), and be tethered to higher-order regulatory functions (a property called composability).
"Much of the regulatory potential of a bacterium is contained in the five-prime untranslated regions (UTRs), which control the expression of physically adjacent downstream genes and have become attractive platforms for a parts-based approach to synthetic biology," Arkin says. "This approach, in which integrated engineered regulatory parts respond to custom inputs by changing the expression of desired genes, must satisfy two criteria if it is to have long-term success. First, the regulatory parts must be easily engineered in a way that yields large homogenous sets of variants that respond to different custom inputs, and second, the parts must be composable such that they can be easily and predictably assembled into useful higher-order functions."
In the five prime UTRs of bacteria, two primary types of regulators can serve as starting points for designing new parts – those that regulate transcriptional elongation, in which cellular inputs are linked to the process by which a sequence of DNA nucleotides is transcribed into a complementary sequence of RNA; and those that regulate translation, in which a ribosome translates the RNA message into a protein. Transcriptional elongation regulators meet the second criterion by featuring versatility and composability that makes them ideal for building custom regulatory functions. Translational regulators meet the first criterion by being easier to engineer and relatively common to all bacteria.
"Our solution for meeting both criteria was to develop an adaptor based on tryptophanase, the catabolic operon for tryptophan that converts regulators of translational initiation into regulators of transcriptional elongation," Arkin says. "Because our adaptor strategy bypasses the otherwise restrictive tradeoff between criterion one and criterion two, we believe it will have a crucial role in the long-term development of five prime UTRs as platforms for the design and integration of custom regulatory parts."
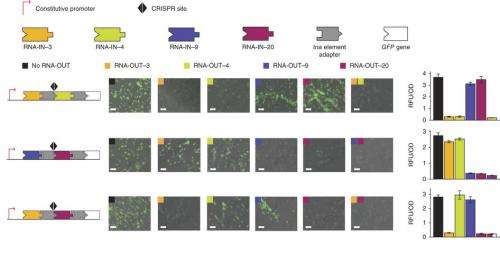
When an E.coli translational regulator was fused to the adaptor created by Arkin and his colleagues, it was also able to control transcriptional elongation. The team applied their adaptor to the construction of several transcriptional elongation regulators that respond to RNA and small-molecule inputs. Included were five mutually orthogonal RNA-triggered attenuators (meaning they can terminate transcription), which the team assembled into logic gates driven by two, three or four RNA inputs that linked to ribosome binding sites. Because their adaptor is so easily linked to ribosome binding sites, a common mechanism in bacteria, the team believes the adaptor will be widely applicable.
"Continued application of our adaptor should produce large collections of transcriptional regulators whose inherent composability can facilitate the predictable engineering of complex synthetic circuits," Arkin says.
The adaptor is ready for use by researchers now, and joins a growing suite of tools created by Arkin's group and their collaborators to make the design and fabrication of microbial systems more reliably predictable.
"Taken together, these tools enable a predictable biological circuit design cycle that dramatically increases the efficiency, safety and sophistication of genetic engineering," Arkin says.
Journal information: Nature Methods
Provided by Lawrence Berkeley National Laboratory





