How E. coli grows its 'nose'
(PhysOrg.com) -- Self-assembling and self-organizing systems are the Holy Grails of nanotechnology, but nature has been producing such systems for millions of years. A team of scientists has taken a unique look at how thousands of bacterial membrane proteins are able to assemble into clusters that direct cell movement to select chemicals in their environment. Their results provide valuable insight into how complex periodic patterns in biological systems can be generated and repaired.
Researchers with Berkeley Lab, the University of California (UC) Berkeley, the Howard Hughes Medical Institute, and Princeton University, used an ultrahigh-precision visible light microscopy technique called PALM - for Photo-Activated Localization Microscopy - to show that the chemotaxis network of signaling proteins in E.coli bacteria is able to spontaneously form from clusters of proteins without being actively distributed or attached to specific locations in cells. This simple organizational mechanism - dubbed “stochastic self-assembly” - is related to the self-organizing patterns first described in 1952 by the British computer scientist Alan Turing.
“It is not widely appreciated that complex periodic patterns can spontaneously emerge from simple mechanisms, but that is probably what is happening here,” said Jan Liphardt, the biophysicist who led this research.
Liphardt holds a joint appointment with Berkeley Lab’s Physical Biosciences Division and UC Berkeley’s Physics Department. He is the principal author of a paper now available on-line in the Public Library of Science entitled: “Self-Organization of the Escherichia coli Chemotaxis Network Imaged with Super-Resolution Light Microscopy.” Co-authoring the paper with Liphardt were Derek Greenfield, Ann McEvoy, Hari Shroff, Gavin Crooks, Ned Wingreen and Eric Betzig.
Key to a cell’s survival is the manner in which its critical components - proteins, lipids, nucleic acids, etc. - are arranged. For cells to thrive, the organization of these components must be optimized for their respective activities and also reproducible for succeeding generations of cells. Eukaryotic cells feature distinct subcellular structures, such as membrane-bound organelles and protein transport systems, whose complex organization is readily apparent. However, there is also complex spatial organization to be found within prokaryotic cells, such as rod-shaped bacteria like E. coli.
“It has remained somewhat mysterious how bacteria are able to organize and spatially segregate their interiors and membranes,” said Liphardt. “Two cells which are biochemically identical can have very different behaviors, depending upon their spatial organization. With new technologies such as PALM, we are able to see exactly how cells are organized and relate spatial organization with biological function.”
PALM and the Chemotaxis Network
In the PALM technique, target proteins are labeled with tags that fluoresce when activated by weak ultraviolet light. By keeping the intensity of this light sufficiently low, researchers can photoactivate individual proteins.
“Since individual proteins are imaged one at a time, we can localize and count them, and then computationally assemble the locations of all proteins into a composite, high-precision image,” said Liphardt. “With other technologies, we have to choose between observing large clusters or observing single proteins. With PALM, we can examine a cell and see single proteins, protein dimers, and so forth, all the way up to large clusters containing thousands of proteins. This enables us to see the relative organization of individual proteins within clusters and at the same time see how clusters are arranged with respect to one-another.”
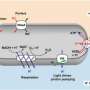
Liphardt and his colleagues applied the PALM technique to the E.coli chemotaxis network of signaling proteins, which direct the movement of the bacteria towards or away from sugars, amino acids, and many other soluble molecules in response to environmental cues. The E.coli chemotaxis network is one of the best-understood of all biological signaling systems and is a model for studying bacterial spatial organization because its components display a nonrandom, periodic distribution in the cell membrane.
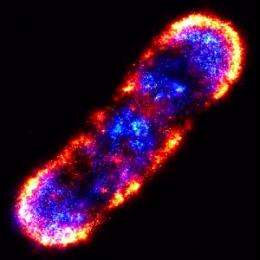
“Chemotaxis proteins cluster into large sensory complexes that localize to the poles of the bacterial cell,” Liphardt said. “We wanted to understand how these clusters form, what controls their size and density, and how the cellular location of clusters is robustly maintained in growing and dividing cells.”
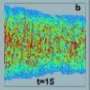
Using PALM, Liphardt and his colleagues mapped the cellular locations of three proteins central to the chemotaxis signaling network - Tar, CheY and CheW - with a mean precision of 15 nanometers. They found that cluster sizes were distributed with no one size being “characteristic.” For example, a third of the Tar proteins were part of smaller lateral clusters and not of the large polar clusters. Analysis of the relative cellular locations of more than one million individual proteins from 326 cells determined that they are not actively distributed or attached to specific locations in cells, as had been hypothesized.
“Instead,” said Liphardt, “random lateral protein diffusion and protein-protein interactions are probably sufficient to generate the observed complex, ordered patterns. This simple stochastic self-assembly mechanism, which can create and maintain periodic structures in biological membranes without direct cytoskeletal involvement or active transport, may prove to be widespread in both prokaryotic and eukaryotic cells.”
Liphardt and his research group are now applying PALM to signaling complexes in eukaryotic membranes to see how widespread is stochastic self-assembly in nature. Given that biological systems are nature’s version of nanotechnology, the demonstration that stochastic self-assembly is capable of organizing thousands of proteins into complex and reproducible patterns holds promise for a wide range of applications in nanotechnology, including the fabrication of nanodevices and the development of nanoelectronic circuits.
Provided by University of California - Berkeley (news : web)