'Disordered' amino acids may really be there to provide wiggle room for signaling protein
Sections of proteins previously thought to be disordered may in fact have an unexpected biological role - providing certain proteins room to move -- according to a study published by researchers at Fox Chase Cancer Center in this month's issue of the journal Structure.
The researchers published the first comprehensive structural study of the protein NHERF1, which serves as a means of bringing together molecular signals between the outer membrane of a cell and the proteins found in the structures that form the cell's cytoskeleton. NHERF1 is representative of a large class of proteins with an array of biological roles, including signaling pathways implicated in diseases such as cystic fibrosis and breast cancer.
"Here we have a molecule that serves an important role in how cells function and survive, but it contains these puzzling 'junk' sequences that don't seem to have any apparent purpose," says the paper's lead author Heinrich Roder, Ph.D., a structural biologist and senior member of the Fox Chase Cancer Center faculty. "Our work suggests that this disorder is really a way of creating flexibility, allowing the protein to function as a molecular switch, a process that is thought to go wrong in certain diseases."
The NHERF1 proved particularly puzzling to researchers as it contains both known structures and large disordered regions. The structured sections of the protein are comprised of two so-called PDZ domains - components well known to scientists as anchors that can help the protein connect to other proteins and cellular structures - and an ezrin-binding motif, a sequence of amino acids that connects NHREF1 to cellular proteins such as ezrin. Together these structures allow NHERF1 to serve as a signaling adaptor, an important link between different proteins in a pathway. Oddly, researchers have noted that the ezrin-binding motif conveniently fits into a cleft on the surface of the PDZ domains, but assumes a helical structure, which is highly unusual for this class of proteins.
As is typical for human proteins, over one third of the amino acids that make up NHERF1 were predicted to be intrinsically disordered, forming no known structures and matching no known evolutionarily-conserved pattern. According to Roder, it is one of these disordered segments, approximately 100 amino acids long, which enable the protein to work. That is, it allows the PDZ binding module to "bite" its own ezrin-binding tail, effectively shutting the protein down.
"The flexibility of the linker and its tendency to be more or less disordered are critical for regulating the balance between internal ("autoinhibitory") forces and external interactions with the protein's signaling partners" Roder says. This idea is reinforced by a recent observation by Zimei Bu, Ph.D., a Fox Chase researcher a coauthor of this study, who found that enzymatic modification (phosphorylation) of amino acids in the disordered region tips the balance towards the more open, active, form of the protein.
"Evolution has provided researchers with convenient modular structures, areas that are repeated over and over again to make up proteins, and so we tend to dismiss the interspersed disordered sequences that don't seem to have any definable structure," Roder says. "Here we show that the weak molecular interactions in a disorganized protein sequence are essential in giving this protein its unique attributes."
It was also this disorganized domain that made it difficult for researchers to create an accurate model of the entire protein, Roder says. Typically, researchers use a technique called x-ray crystallography, in which they can tell a protein's structure from how crystals made from the protein samples scatter x-rays. The disorganized section of NHREF1 makes it nearly impossible to create the necessary crystals to use this technique. Instead, Roder and colleagues used a technique called nuclear magnetic resonance spectroscopy, which can determine a protein's shape by measuring how the individual atomic nuclei of a protein interact with an intense magnetic field.
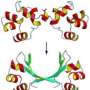
Since NMR spectroscopy works best with small proteins - not large flexible molecules like NHREF1 - Roder, his staff scientist Hong Cheng, Ph.D., and their colleagues came up with an innovative technique that allowed them to look at the protein from numerous angles with the NMR spectrometer, and subtract out the overlapping areas to form a more complete picture of the molecule. This picture enabled them to see what NHREF1 looks like in both its "on" and "off" conformation.
"When it is not working, the protein is in this 'off' conformation until acted upon by outside agents" Roder says. "It is a way for the cell to shut off a signaling pathway when it is not in use."
According to Roder, these findings may provide researchers a new way of looking at proteins like NHERF1 and their physiological role in cells.
Source: Fox Chase Cancer Center (news : web)