Stir no more: Scientists show that draining speeds up bioassays
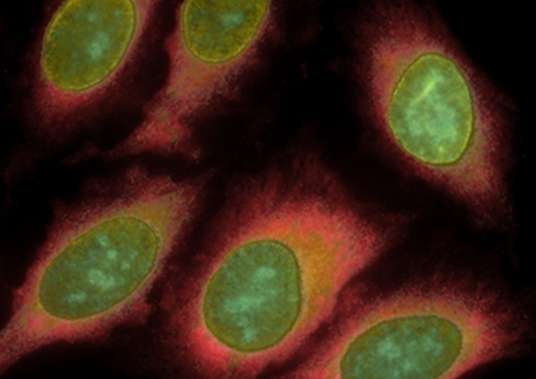
For many research scientists, idle time has long been an unwelcome feature of the discovery process. Advances in cellular biology have yielded popular and powerful tools to detect cellular proteins and DNA—largely by exploiting the unique and intricate interactions between these microscopic molecules. Scientists use these tools to advance research and diagnose diseases. But these come at a cost in time—from hours to days—before they yield accurate answers.
Three scientists at the University of Washington have proposed a way to speed up this waiting game. Their solution, reminiscent of the magic behind washing machines, could reduce wait times to a fraction of what they once were. As they report Jan. 8 in the journal Small, biological assays that once took hours could instead take minutes.
"These are very common assays," said Xiaohu Gao, a UW professor of bioengineering and senior author on the paper. "Most scientists were willing to wait hours and hours because they had no choice."
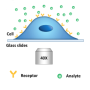
Many of today's biological assays use molecules such as antibodies to detect specific types of cellular proteins or pieces of DNA. These "detector molecules" only bind to specific targets, such as a certain class of cellular proteins, and include additional components such as nanoparticles or dye molecules to emit light if they successfully bind. These assays have revealed where different proteins are found in cells and helped diagnose diseases.
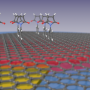
But these tests take hours or days to complete. The detector molecules, suspended in a fluid, float around while their targets—whether cellular proteins or pieces of DNA—are adhered to the hard, flat surface of a small plate or petri dish. While bulky detector molecules close to the surface can easily find and bind to their targets, molecules further up in the fluid column move slowly due to their size. It can take hours for enough detector molecules to diffuse down and bind to their targets to produce a visible color change.
"We call this 'diffusion limitation,' and it's a serious problem since both the antibody and nanoparticles are so large," said Gao. "People have proposed solutions—like stirring or gently rocking a reaction plate to mix the solution. But when we tested this we saw that stirring and rocking only improved the reaction time by 3 to 5 percent. That's not enough."
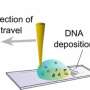
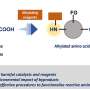
Gao and his team were prompted to tackle the problem of diffusion limitation after they developed a new staining assay but its long reaction times made their protocol impractical. Inspired by studies of fluid dynamics, they decided to work around the problem of diffusion limitation. Instead of waiting for detector molecules to drift down to the surface of the plate, they simply allowed detector molecules close to the surface to bind. Then, they drained the solution from the plate, mixed it, put it back on the plate and repeated this cycle dozens of times—which they call cyclic solution draining and replenishing.
"In a washing machine, you squeeze water out and put it back in," said Gao. "Dry and re-soak. Dry and re-soak. This is exactly the same mechanism: Drain the fluid completely and then put it back on the plate. That's much more efficient than simply stirring it around."
To drain fluid from the plate, they covered the plate with a seal and inverted it. To "re-soak," they flipped the plate upright again. The flipping action helped mix the detector molecules in the fluid, which sped up the total reaction time.
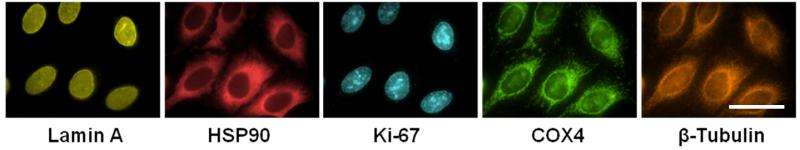
They tested cyclic solution draining and replenishing with two types of antibody staining techniques, ELISA and immunofluorescence microscopy. Reaction times for both were cut substantially with this drain and re-soak approach. In one case, what was once a one-hour incubation time was cut to just seven minutes. Though sealing and flipping the plate, which they accomplished mechanically, might be impractical for other tests, there are other ways to "drain" a plate.
"We used gravity because we wanted to show that draining would work," said Gao. "But you could use air bubbles or centrifugation to drain as well, for example. There are lots of possibilities."
If so, this simple work-around for the problem of diffusion limitation could slash waiting times for experiments. This would also impact other fields, reducing the wait times for medical test results to come back or speed up chemical engineering protocols.
"Really, this was a common problem that no one before had made a link to. But here we have, and it's so simple," said Gao. "When we prepare tea, we don't let it sit there or shake the cup. We repeatedly lift, drain the tea bag, then lower it into the hot water. That's what we've done here."
More information: Eliminating Size-Associated Diffusion Constraints for Rapid On-Surface Bioassays with Nanoparticle Probes, DOI: 10.1002/smll.201503101 , onlinelibrary.wiley.com/doi/10 … /smll.201503101/full
Journal information: Small
Provided by University of Washington