Bioinformaticians make the most efficient search engine for molecular structures available online
"CSI: Crime Scene Investigation" is a well-known American TV series in which murder cases are solved with the help of precise forensic science. Although Prof. Sebastian Böcker and his team at the Friedrich Schiller University in Jena, Germany, have nothing to do with CSI, these bioinformaticians are experienced readers of trails. They hunt for molecular structures of metabolites, which are chemical compounds that determine the metabolism of organisms. "Metabolites can provide detailed information about the state of living cells, provided that researchers are successful in identifying and quantifying the multitude of metabolites," Prof. Böcker explains.
This process is highly complex and seldom leads to conclusive results. However, the work of scientists all over the world who are engaged in this kind of fundamental research has now been made much easier: The bioinformatics team led by Prof. Böcker in Jena, together with their collaborators from the Aalto-University in Espoo, Finland, have developed a search engine that significantly simplifies the identification of molecular structures of metabolites. In the newly published edition of the well-known science magazine Proceedings of the National Academy of Sciences (PNAS), they present their search engine 'CSI:FingerID.'
In this case, CSI stands for "Compound Structure Identification," and is based on combining a variety of methods. To begin with, metabolite samples to be analysed undergo a so-called tandem mass spectrometry run. "During this step, molecules are dismantled into smaller fragments and their molecular weights are identified," Böcker explains.
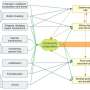
The resulting spectra give information about the chemical composition of metabolites, but this information is not yet adequate to draw conclusions about the molecular structure. This is where the newly developed search engine comes into play. It works in a similar way to an internet search engine, but instead of searching for keywords, the tool looks for molecular information that translates the given mass spectrum into a structural formula.
After the mass spectrum has been submitted to the search engine, 'CSI:FingerID' trawls a number of online molecular structure databases, where scientists throughout the world publish information and structural formulae of both newly discovered and long-known metabolites. A single 'CSI:FingerID' search results in a list of possible candidate structures which best correspond to the spectrum.
Reduce the Number of Possible Compounds
"After obtaining the list of possible candidates we still don't know with absolute certainty which metabolite we are dealing with. But when we can reduce the number of possible compounds from several thousand down to perhaps ten, then this is huge progress," says Böcker. "Because precise lab tests to identify compounds can be expensive and time-consuming, so distinguishing among thousands of possibilities is usually impossible – but testing just ten compounds is often feasible." And, as the relevant databases also grow constantly – with an average of ten entries being added per minute on a worldwide basis – the search results become consistently more precise.
The bioinformaticians show in this new study that they obtain a significantly higher hit ratio with their method than any other method that has been used so far. To this end, they have validated their search engine with more than 6,000 test substances. As well as using 'CSI:FingerID' themselves to analyse naturally occurring metabolites, Prof. Böcker and his Jena team have made the search engine freely available to the international scientific community.
More information: "Searching molecular structure databases with tandem mass spectra using CSI:FingerID," PNAS 2015, DOI: 10.1073/pnas.1509788112
Journal information: Proceedings of the National Academy of Sciences
Provided by Friedrich Schiller University of Jena