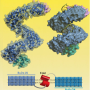
Head formation of clawed frog embryos

On July 9, 2014, Dr. Yuuri Yasuoka in the Okinawa Institute of Science and Technology Graduate University's Marine Genomics Unit, published a research paper explaining a key mechanism in formation of the head in frogs. The work was done in collaboration with researchers at the University of Tokyo. Previous studies had reported genes involved in head development. However, it still remained unclear how those genes interact with each other for head formation as a whole. By employing Next-Generation sequencing techniques, which provide scientists with massive amounts of DNA sequence data, Dr. Yasuoka has uncovered a genetic mechanism underlying head formation, which is one of the most important processes in animal development.
For centuries, scientists have explored how fertilized eggs develop into mature organisms. Since DNA in the nucleus of a cell was discovered as the substance of genetic inheritance, genetic studies have made huge progress. In the 21st century, advances in genome analysis led to decoding of human genome, an achievement that represents a quantum leap in human self-knowledge. Today, in addition to asking what kind of genes are functioning in a specific part of the body, scientists are also asking how individual genes influence each other to organize the body.
In this study, Dr. Yasuoka prepared the so-called "head organizer cocktail" comprised of several proteins required for head formation during early development of a fertilized egg and introduced it into a frog embryo. As a result, an additional head was formed on the ventral side, or the stomach side, of the embryo, indicating that those proteins induce tissues to form a head. Dr. Yasuoka also demonstrated that an embryo that lacked those genes could not form a head properly. He then tried to discover where those proteins are located in the genome and how they affect other genes for head formation. By using Next-Generation sequencers, Dr. Yasuoka decoded frog DNA sequences bound by those proteins and successfully created a genetic map of protein-binding regions in the genome in a process called mapping. These proteins, generally referred to as "transcription factors," are known for their role in enhancing or inhibiting the activity of other genes. This study made it possible to evaluate how these proteins enhance and inhibit genes for head formation by delving into transcription factor binding regions in the genome, known as cis-regulatory modules. This study provided detailed insight into the regulatory mechanisms of gene expression.
Changes in developmental processes in individual organisms lead to evolution over a long period of time. "Evolution continues forever and our understanding of it also forever remains within the boundary of speculation," Dr. Yasuoka said. "What we can do is to collect as much information as possible about organisms for use in experiments and make the most use of scientific knowledge to fill in gaps in the evolutionary history of life."
More information: "Occupancy of tissue-specific cis-regulatory modules by Otx2 and TLE/Groucho for embryonic head specification." Yuuri Yasuoka, et al. Nature Communications 5, Article number: 4322 DOI: 10.1038/ncomms5322.
Journal information: Nature Communications
Provided by Okinawa Institute of Science and Technology