Split-second snapshots of protein development

The birth of a protein is one of the most fundamental aspects of life as we know it, yet, surprisingly, there is still a lot that scientists do not know about them.
Now split-second snapshots of this vital and mysterious process, developed by Samuel Cho, an assistant professor of physics and computer science, and senior Jessica Leuchter ('14), could someday lead to more effective antibiotics.
The birth of a protein
Proteins are the worker bees of cells. They get rid of waste, transmit cellular signals and carry out the chemical reactions that enable the human body to function. Without proteins, cells would be unable to function, replicate, and die.
This is not always a bad thing though, said Leuchter, a chemistry major who has been conducting research with Cho for two years. Viruses, bacteria and cancer cells need proteins to reproduce like everything else living in the human body.
Considering a 2013 CDC report noted that more than 2 million people get sick and roughly 23,000 people die each year as a result of drug-resistant bacterial infections, a new drug that could specifically target and halt protein formation in these malignant organisms would be a powerful tool for combatting diseases.
The problem with creating such a drug is scientists know very little about the microscopic environmental conditions and structures involved in creating proteins. This lack of awareness makes it difficult to identify targets inside a cell that drugs can be designed to attack.
Cho explained this is mostly due to the fact protein formation takes place at an incredibly small scale.
"Imagine the smallest thing you can think of and this is probably a lot smaller than that," Cho said. "The entire process occurs in a space 100 billion trillionth the size of a water droplet. Even the best microscopes we have are not powerful enough to study this in the lab."
Taking a closer look
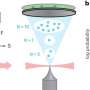
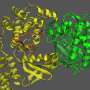
This is where Cho and Leuchter's work comes in. They created a computer model that simulates a required protein called Methionyl-tRNA Synthetase at one of the very first stages of protein formation in an E. coli bacteria cell. Their model enables them to accurately simulate processes at the molecular scale and identify and characterize previously unknown structures and crucial details of the chemical reactions that can then be used as drug targets. Then, an experimentalist can verify the targets Cho and Leuchter identify as critical in the lab.
"Thankfully we have a collaborator in the chemistry department who is the foremost expert on Methionyl-tRNA Synthetase, Rebecca Alexander," Cho said. "It's really nice having the authority on the subject in the next building so that she can say yes we have identified a new target or no we haven't."
New ways to fight disease
Cho said the next step in the research will be to tweak the computer model so it can be used to examine other proteins.
"There are 20 other proteins similar to the one our model is designed to analyze," Cho said. "Hopefully by learning about this one we can apply some of the same approaches to other proteins."
Leuchter said figuring out how to disrupt the formation of different proteins via computer models is an important step towards coming up with new treatments for cancer.
"Our model could be fundamental in finding new pharmaceutical targets," she said. "It could also stop a defective protein from forming."
In addition, models like theirs can be used to develop new antibiotics to kill drug-resistant bacteria or "super-bugs."
"What if we could turn important reactions off in the bacteria?" Cho asked. "A clearer picture of what is actually going on inside a cell will help us figure out what is an important interaction in terms of making proteins and what isn't."
Looking ahead to the future
Earlier this month, Leuchter traveled to Germany, where she presented a poster on her computer simulation at the 552nd WE-Heraeus Seminar at the Karlsruhe Institute of Technologie. The conference, one of the most renowned meetings for biophysicists in the world, featured global experts presenting on progress in the interdisciplinary field.
After graduating this spring, Leuchter says she plans to take a year off before coming back to school to pursue a MD-PhD.
Looking back, she said working with Cho has been the highlight of her Wake Forest experience.
"Dr. Cho is such a good mentor. He is concerned not just with how I do in class but also with how I am doing socially and mentally," she said. "He has helped to round out my skills and made me realize I'd like to pursue a career coming up with new ways to fight different kinds of disease."
Provided by Wake Forest University