Scientists combine bacteria with liquid crystals

(Phys.org) —When swimming around, bacteria aren't good with the "pool rules." In small quantities, they'll follow the lanes, but put enough together and they'll begin to create their own flow.
In a collaboration between the U.S. Department of Energy's Argonne National Laboratory and the Liquid Crystal Institute at Kent State University, researchers combined bacteria with liquid crystals and observed how the bacteria swam around and interacted with the medium, forming "living liquid crystals" that may have interesting applications for new material design and fabrication.
Liquid crystals are intermediate materials that exhibit some of the behaviors of a liquid and some of those of a crystalline solid. For example, a liquid crystal has a liquid-like flow, but on the molecular level it may appear more crystalline.
When they are dissolved in water, the molecules that make up a liquid crystal tend to form long rod-like structures that prefer to organize along one dimension, called a director. The director is symmetrical—it does not have poles along its axis.
Using a solution known as "Terrific Broth" – so known for its usefulness as a bacterial growth medium – the researchers cultured a colony of Bacillus subtilis bacteria and transferred it to a liquid crystal.
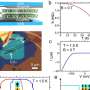
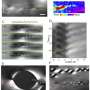
When the researchers looked at the bacteria in the liquid crystal using a powerful optical microscope, they saw that they moved in a surprising way. Each B. subtilis bacterium has a long, thin tail called a flagellum, which the bacterium spins like a screw to propel itself. Although the bacteria originally orient themselves parallel to the director, the spinning of the flagella as the bacteria swim locally changes the director's alignment into a wave-like pattern.
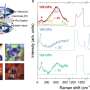
As the researchers increased the density of bacteria in the liquid crystal solution, they saw a second surprising effect. Because of the collective effects, the swimming bacteria began to create "stripes" of different director orientations within the liquid crystal. These stripes could then be erased by depriving the bacteria of oxygen.
"The system is extremely sensitive," said Argonne materials scientist Andrey Sokolov, one of the authors of the study. "There is always an interplay between the bacterial forces and liquid crystal forces, and the behavior of the material depends on which one wins in the end."
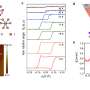
According to Argonne materials scientist Igor Aronson, the study's lead author, it only takes a bacterial concentration of 0.2% for the bacteria's collective swimming to begin to overtake the natural order of the liquid crystal.
The researchers hope that eventually studies on living liquid crystals as well as other similar "living materials" will pay dividends for a number of different applications. "Our principal goal is understanding active materials," Aronson said. "We want to see how we can design new ways to consume energy from the environment and use it for self-assembly and self-repair."
The scientists also noted that their method may have other implications beyond the living liquid crystals themselves. Because the wake created by the oscillations of the flagella in the liquid crystal is so much larger than the nanometer-wide flagella themselves, the researchers were able to observe the action of the flagella with an optical microscope. "The flagella are only a few tens of nanometers in diameter, and in the past we've had to use electron microscopes in order to see it," Aronson said.
"This could have bigger implications in terms of ways to visualize other nanoscale objects," Sokolov added.
Funding for the study was provided by DOE's Office of Science and the National Science Foundation, and results appeared as a paper titled "Living liquid crystals" in the Proceedings of the National Academy of Sciences.
More information: Shuang Zhou, Andrey Sokolov, Oleg D. Lavrentovich, and Igor S. Aranson. "Living liquid crystals." PNAS 2014 111 (4) 1265-1270; published ahead of print January 13, 2014, DOI: 10.1073/pnas.1321926111
Journal information: Proceedings of the National Academy of Sciences
Provided by Argonne National Laboratory