Amending the Tree of Life: Article advocates for shift in evolutionary and genomic research
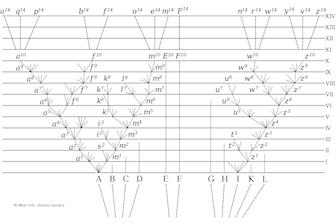
Charles Darwin famously included only one illustration in his book "On the Origin of Species" published in 1859. It was a diagram of how species originate through time in a manner that, on paper, resembles a family tree. Since then, geneticists have been piecing together relationships among life's many creatures by DNA comparison to produce what is called the Tree of Life: a tree-shaped diagram of how everything on Earth is related. This type of diagram has dominated the perspective of evolutionary biologists for more than 150 years.
More than a decade ago, Scot Kelchner, associate professor of systematics and evolution in Idaho State University Department of Biological Sciences, began using a different and less popular mathematical approach for displaying evolutionary history in his research. Instead of simple tree diagrams to explain the inter-connections of life, he used networks that allowed complex genetic relationships to be shown and tested.
These "phylogenetic networks" are now the subject of his August 2013 cover article in Trends in Genetics, a prominent and highly respected scientific journal followed closely by geneticists around the world. Kelchner is a communicating author on the article, which he helped to organize and write. His coauthors are eminent mathematicians, computational scientists, and biologists from seven countries around the world.
The research team formed in October 2012 in Leiden, Netherlands, during a weeklong "think tank" on phylogenetic networks at which Kelchner gave an invited address. His visit was sponsored in part by the ISU Office of Research.
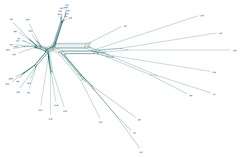
It was at this meeting that Kelchner and his colleagues decided to write a prominent paper showing a united front among scientists and mathematicians on the importance of networks for the future of evolutionary biology and genome research. The team's aim is to make network use as routine and easy as tree use is now for describing pathways of evolution.
Moving from tree-like depictions of evolution to network diagrams is an effective way to amend the Tree of Life without dismissing it, as some authors have controversially recommended says Kelchner. He notes that because networks allow researchers to show and measure more complexity in evolution than a simple tree shape can represent, they open a new door to the way scientists think about and study the evolutionary history of genes, genomes and organisms.
The tools for network analysis have been around for more than a decade, but biologists have been slow to adopt them. This peer-reviewed cover article in Trends in Genetics, commissioned by the editors, is a clear signal to the world's geneticists that network approaches are the next major advance in the study of genome evolution.
More information: www.cell.com/trends/genetics/issue?pii=S0168-9525%2813%29X0008-3
Journal information: Trends in Genetics
Provided by Idaho State University