A roundabout route to protein production
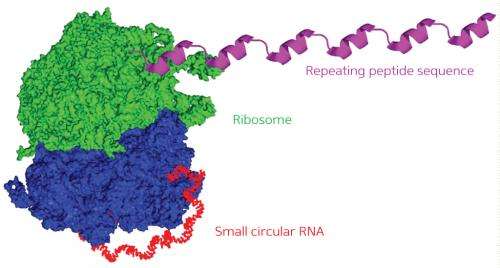
Proteins are typically encoded by linear strands of messenger RNA (mRNA). These mRNA molecules are translated into polypeptide chains by ribosomes, with each ribosomal read-through of the mRNA generating a single, discrete copy of the encoded protein. New work from Hiroshi Abe and Yoshihiro Ito of the RIKEN Nano Medical Engineering Laboratory now demonstrates the potential to turn a closed loop of mRNA into a steady assembly line for continuous protein production.
Scientists routinely employ a technique called 'rolling-circle amplification' to generate many continuous copies of a single circular DNA template, and Abe had become interested in devising an equivalent approach for protein production. "I came up with the idea of using circular RNAs for translation," he says. Other researchers have attempted this in the past in the bacterium Escherichia coli, but came to the conclusion that the process was considerably less efficient than for linear RNA templates.
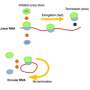
Abe and Ito therefore set out to systematically test different constructs in order to identify factors that could boost or undermine the productivity of circular RNAs in E. coli cellular extracts. A typical protein-coding RNA begins with a ribosomal binding sequence and a 'start codon' and ends with a 'stop codon' that causes the ribosome to detach from the finished polypeptide chain. The researchers built four different circular RNAs of different lengths, featuring 'infinite' coding regions that lack stop codons, and found that all but the smallest of these could successfully generate continuous chains of tandem protein repeats (Fig. 1).
Subsequent experiments permitted a direct comparison of productivity between a 126-nucleotide circular RNA and its linear equivalent. Abe was surprised to observe that the circular molecule exhibited far superior performance, generating 100-fold more protein than the linear template. He hypothesizes that this efficiency boost arises from the fact that linear templates require ribosome 'recycling', in which ribosomes released at the end of translation must subsequently seek out new binding sites, while closed templates keep the same ribosome locked in on a continuous circular path. "We have proved that the rate-limiting step of protein translation is ribosome turnover," says Abe.
Circular RNAs encoding fluorescent proteins could offer useful 'tags' for labeling molecular targets in biochemical assays, but Abe notes that this rolling-circle protein production strategy might also provide an effective tool for churning out long strands of protein for biotechnology or materials applications. "It could be used for the production of spider silk, collagen, or other proteins with repeated sequences," he says.
More information: Abe, N., et al. Rolling circle amplification in a prokaryotic translation system using small circular RNA. Angewandte Chemie International Edition 52, 7004–7008 (2013). dx.doi.org/10.1002/anie.201302044
Journal information: Angewandte Chemie International Edition
Provided by RIKEN