New methodology for the analysis of proteins
A study led by the professor of Biochemistry and Molecular Biology from the Faculty of Chemistry of the UB Modesto Orozco, and by Xavier Salvatella, from the Department of Biochemistry, both ICREA scientists at the Institute for Research in Biomedicine (IRB Barcelona), has devised a new strategy to study the shape of proteins.
According to Orozco, "by combining computational modeling and experimental physicochemical techniques, we have revealed the structures of proteins, which, until now, were unachievable because of technical barriers". Results have been published on the journal Proceedings of the National Academy of Sciences (PNAS).
The research, carried out within the joint programme IRB Barcelona - Barcelona Supercomputing Center (BSC) —centres located at the BKC—, represents an advance in protein structure research. Michela Candotti, the first author of of the paper, states that "to know the shape that proteins have is essential to perform any analysis. A wire can be a paperclip, a staple or a spring, depending how it is folded". This remark is especially relevant given the multi-functional nature of many proteins.
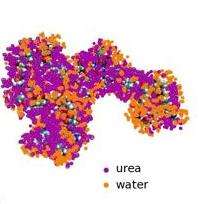
In the study researchers have been able to describe the chemical mechanisms by which compounds such as urea unfold proteins. "This was a debate that started in the 60s and now, with this work, it can be considered closed", explains Orozco. Furthermore, they have established a new strategy that will allow them to decipher the conformation of the Intrinsically Disordered Proteins (IDP). IDPs are a group of proteins without a rigid structure that comprise a large part of the proteome; however, little is known about them. "Our results will contribute to research into diseases that involve IDPs, such as cancer, Parkinson's or Alzheimer", affirms Salvatella. Finally, scientists have identified the first steps in protein folding, another aspect which is discussed at great lenght.
More information: Candotti, M. et al. Towards an atomistic description of the urea-denatured state of proteins. Proceedings of the National Academy of Sciences (PNAS), (early edition) 25th March 2013. DOI: 10.1073/pnas.1216589110
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Barcelona