A new computational method for timing the tree of life

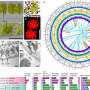
With its deeply embedded roots, sturdy trunk and dense profusion of branches, the Tree of Life is a structure of nearly unfathomable complexity and beauty. While major strides have been made to establish the evolutionary hierarchy encompassing every living species, the project is still in its infancy.
At Arizona State University's Biodesign Institute, Sudhir Kumar has been filling in the Tree of Life by developing sophisticated methods and bioinformatics tools. His latest research, which appeared on the advance online edition of the Proceedings of the National Academy of Sciences will uniquely enable scientists to analyze very large datasests to set time to the multitude of branching points (nodes) on the tree, each representing a point of species divergence from a common ancestor. The new method differs significantly from currently used techniques and excels in providing results of equal or greater accuracy at speeds of 1000 times or faster.
For the proper study of evolutionary history, two components are key: the relationships between organisms (known as phylogeny) and their times of divergence. As Kumar explains, the powerful technique for estimating the time of divergence between species was initially realized over four decades ago, when the concept of molecular clocks was introduced. Initially the idea rested on the assumption that alterations in either the amino acid sequences of proteins or the nucleotide sequences of DNA between various species accumulate at a uniform rate over time and can be used to evaluate divergence times. The resulting phylogenetic structure is known as a "TimeTree," that is, a tree of life scaled to time.
Prior to the use of molecular clocks, morphological changes between species were the primary means of identifying divergence times. Since then, molecular clocks have proved a vital tool for evolutionary biologists, supplementing the fossil record and providing a powerful means to time the divergence of species.
But there is a complication. The rate of change measured by molecular clocks can vary—sometimes radically—between groups of species. Rather than an ordered world running on a universal clock time, the Tree of Life is more like an antiques shop where clocks run at different speeds in different species.
Many approaches for dealing with this conundrum have been applied successfully, but their complexity rises exponentially with the number of species involved. Often such calculations swallow vast amounts of computing time, even for data sets of modest size.
By contrast, the new simplified method (known as RelTime) produces rapid results. Its main purpose is to estimate relative times of divergence. This avoids the need to use the fossil record, which is otherswise required in order to obtain absolute times.
"If, for example, we can establish that human and chimp divergence is 5 times younger than the human and monkey divergence, that would be very useful," Kumar says. "What our method can do is to generate such relative time information for every divergence in the Tree of Life—without using the fossil record or other complicated model parameters. " Once relative times for all the nodes on the tree of life are established, fossil calibration points for which a high degree of confidence exists can be applied post hoc to add the absolute time dimension.
Kumar points out that rapid DNA sequencing has allowed for huge datasets of comparative molecular sequences to be generated. Analyses of even a few hundred sequences through current methods however can severely strain computer resources and more massive data sets now being generated can not be solved in reasonable time through current methods, so a fresh approach was needed.
Using RelTime and restricting the analysis to relative divergence times produces results for large phylogenetic trees in hours rather than days. It can also deliver better accuracy, particularly when datasets are enormous and species of interest are from vastly different groups.
"The uses of such technique are only limited by one's imagination. They can be used to estimate the origin of familiar species, emergence of human pathogens, and so forth," Kumar says. "The method is applicable wherever you work with sequences and trees."
RelTime may also help sort out troubling disparities between divergence times based on the fossil record versus those established through the use of molecular data. Examples of dramatic discrepancies between fossils and sequence change measurements have provoked spirited debate, particularly concerning the adaptive radiation of mammals posited to have occurred at the time of dinosaur extinction some 65 million years ago and the divergence of specific animal phyla believed to date to the beginning of the Cambrian period (~500 Mya). In both cases, for example, the molecular dates are about 50 percent older than fossil dates.
The ongoing Timetree of Life project will have important ramifications for many fields of research, providing deep insights into comparative biology, as well as generating data of relevance for paleontologists, geologists, geochemists, and climatologists. Establishing a comparative biological timeline synchronized with Earth history will enable scientists working in diverse areas to explore the long-term development of the biosphere and investigate the evolutionary underpinnings of all life.
In addition to his appointment at the Biodesign Institute, Sudhir Kumar is a professor in the College of Liberal Arts and Sciences, School of Life Sciences
Journal information: Proceedings of the National Academy of Sciences
Provided by Arizona State University