Keeping up with embryogenesis: New microscope lets users track individual cells as they move, divide

(Medical Xpress) -- The transformation of a fertilized egg into a functioning animal requires thousands of cell divisions and intricate rearrangements of those cells. That process is captured with unprecedented speed and precision by a new imaging technology developed at the Howard Hughes Medical Institute’s Janelia Farm Research Campus, which lets users track each cell in an embryo as it takes shape over hours or days.
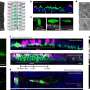
In a single movie – a compilation of a million images captured over about 20 hours – viewers can see biological structures begin to emerge as a simple cluster of cells morphs into an elongated body with tens of thousands of densely packed cells. The movie concludes when the embryo begins to twitch, driven by contractions of its newly formed muscles—moments before the hatching of a fruit fly larva just half a millimeter long.
The imaging technology – driven by a high-speed, non-invasive light microscope that simultaneously captures images from four different angles – is described in the June 3, 2012, issue of the journal Nature Methods. According to its inventor, Janelia Farm fellow Philipp Keller, the new technology will enable quantitative analyses of developmental processes that were not possible with previous imaging methods. “This is not just a quantitative improvement [over previous technologies],” Keller says. “It is the difference between being able to quantitatively measure this process and not even being able to follow the cells.”
Like many developmental biologists, Keller, who works at the interface of biology, physics, and computer science, wants to understand how a single cell transforms into a complex organism. “It’s a really complex process, and if you want to understand it, I think it is really important that you are able to visualize and observe it,” he says. But conventional light microscopes are too slow to follow the rapid cellular migrations that occur at this stage of life, and also tend to damage a living embryo with light before its development is complete. So scientists have been forced to piece together snapshots of different stages of development from multiple organisms, and make assumptions about the changes that must have occurred in between what they actually see.
“This approach is incompatible with the highly dynamic process of development,” Keller says. “Snapshots just don’t capture the essence of it.” Furthermore, cells may divide and rearrange themselves differently each time development occurs, so combining observations from multiple organisms could be misleading.
To better follow development, Keller says, scientists needed a way to image at high speeds, over long periods of time, without interfering with the processes they wanted to observe. Frustrated by the limitations of existing technology, Keller set out to design a microscope that would let him investigate the biological questions that fascinated him most.
The new microscope, which Keller’s team calls the SiMView light sheet microscope, builds on technology Keller developed with colleagues at the European Molecular Biology Laboratory in Heidelberg, Germany to address one of the concerns associated with conventional light microscopy: the damaging effects of exposing a sample to light. Light is essential to activate and visualize the fluorescent labels researchers attach to molecules inside cells, but too much of it damages cells and causes fluorescence to fade. Keller and his colleagues reduced this damage by illuminating only a thin section of a sample at a time with a scanned sheet of laser light, while a detector records the part of the sample being lit up.
The light sheet technique was fast, and could capture the rapid process of development in samples that were not too thick or opaque. In larger or less translucent embryos, however, light from the microscope would scatter as it passed through layers of cells, blurring the image. In a fruit fly embryo, for example, only about 30 percent would be visible.
“You really lose a lot of the information,” Keller says. “If you want to study cellular dynamics during development to get a complete, dynamic building plan of the entire animal, you can’t afford to not see parts of the sample.”
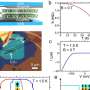
The solution was, in concept, simple: build a light sheet microscope that collects images from multiple views at the same time. “Instead of looking at the sample from one direction, we look at it from four directions at the same time,” Keller explains.
In the new microscope, two light sheets illuminate a sample from opposite directions, and two detectors collect the resulting fluorescence. Each combination of illumination and detection lens provides a different perspective.
Capturing those four views simultaneously eliminates delays caused by rotating a sample repeatedly into new positions – a traditional approach to multi-view imaging – and makes the microscope very fast. “We tried to push the speed as much as we could,” Keller says. This version is as much as 50 times faster than light sheet microscopes that capture multiple views sequentially, he says.
Coordinating the microscope’s components to illuminate the sample and detect fluorescence at the right time and place requires tremendous precision: even fluctuations in room temperature during the course of an experiment can shift components out of alignment. Timing is equally important: light sheets that cross paths can blur an image, so they must be staggered by a few milliseconds. To maintain precision, the SiMView is equipped with an electronics system that makes adjustments in real time.
Once Raju Tomer, a researcher in the lab, and Keller had implemented the design of the microscope, the team had to confront another hurdle: managing and interpreting the tremendous amount of data it generated. The default setting for the microscope, Keller says, is to collect 350 megabytes of data per second. Imaging a single sample over a full day can produce up to several tens of terabytes of data -- about the amount of data stored in the complete Library of Congress archive. For experiments designed to compare development under different genetics or conditions, many times that amount will need to be stored and analyzed. “We want the data,” Keller says. “It’s not a problem, but it is a challenge.”
Anticipating that challenge, lab members Fernando Amat and Khaled Khairy devoted their attention to developing computational methods to identify and track individual cells in the movies produced by the microscope. Those methods are an essential part of what Keller calls a complete technology framework for imaging living samples. The team is now collaborating with Janelia Farm colleagues Kristin Branson and Eugene Myers to develop new computational strategies to automate data analysis,
Having focused on developing that technology for the past two years, Keller says he is looking forward to shifting more of his attention to biology. He anticipates that his lab will continue adapting the microscope and making its computational processing more efficient and powerful. “But we now have the technology to answer most of the questions we are interested in,” he says.
Included in the Nature Methods publication is a movie showing the complete development of the nervous system in a fruit fly embryo. Keller explains that individual cells can be followed as they give rise to sensory organs, brain lobes, and other structures. At the same time, the microscope offers resolution high enough to zoom in and see the fast dynamic processes at the tip of a neuron’s axon.
Anticipating future experiments, Keller says the microscope will let his team not only trace cell lineages in the embryo, but begin to manipulate development to probe the mechanisms that control it. With Janelia colleagues, he has begun to explore other possibilities, tracking development in larger, more complex samples, such as mouse embryos and fruit fly larvae, and monitoring signaling in the zebrafish brain.
“The microscope isn’t the limiting factor anymore,” Keller says. “That’s really exciting.”
More information: DOI: 10.1038/nmeth.2062 , DOI: 10.1038/nmeth.2064
Journal information: Nature Methods
Provided by Howard Hughes Medical Institute