X-rays help advance the battle against heart disease
(PhysOrg.com) -- Scientists from Imperial College London and Diamond Light Source have revealed the structure of a cholesterol-lowering-drug target. Published in the journal Nature, this finding could lead to much more effective drugs to tackle high cholesterol levels, a condition that increases the risk of heart disease.
The researchers from Imperial College London used intense X-rays, generated by the Diamond synchrotron and the European Synchrotron Radiation Facility (ESRF), to determine for the first time the structure of bacterial homologue of the Apical Sodium dependent Bile Acid Transporter (ASBT) protein, a target for hypercholesterolemia drugs since it can affect the level of cholesterol in the blood.
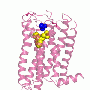
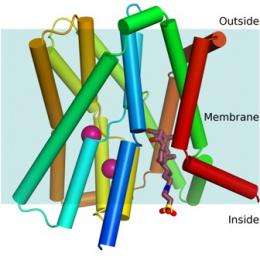
Picture to the right shows a cartoon representation of the ASBTnm structure embedded in the membrane. The protein transports bile acids across the membrane. A bile acid has been trapped in a cavity on the inside face of the protein (shown in wine-red). Energy to drive the transport is provided by sodium ions. Two sodium ions are bound to the structure and these are shown as pink spheres.
In the liver, cholesterol makes bile acids which are used in the intestine to absorb fat. These bile acids are then reabsorbed by ASBT to be transported to the liver and recycled. It is known that by blocking ASBT, bile acid levels returning to the liver are lowered, the liver therefore converts more cholesterol into bile acids, which lowers the level of cholesterol in the blood.
“There are currently a number of existing ASBT inhibitors effective in animal models, which were developed without structural knowledge of the protein. Now that we know the shape and size of the drug-binding site within a bacterial model of the protein, this detailed structural information should enable the design of improved drugs which are much more targeted and will “fit” much better.” said Professor So Iwata, David Blow Chair of Biophysics at Imperial College London.
This new knowledge could have a wider impact on drug design. Dr Alexander Cameron from Imperial College London and the Membrane Protein Laboratory at Diamond explains: “As some drugs are poorly absorbed in the intestine or need to be targeted to the liver, ASBT has also received attention as a pro-drug carrier, capable of transporting various compounds coupled to bile acid. This means that there could be scope to improve a number of drugs tackling different problems, for example, cytostatic compounds targeting liver tumours.”
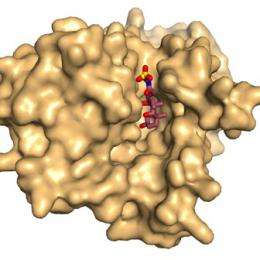
Picture to the left shows a surface representation of ASBTnm looking from the inside face of the membrane showing bile acid bound in a deep cavity.
ASBT is a membrane protein, one of over 7,000 within the human body, of which many are important drug targets. Over 50% of current commercially available drugs target membrane proteins but they are notoriously hard to crystallise – a step that is a pre-requisite in solving protein structures using a synchrotron. Dr David Drew, Royal Society Research Fellow in the Life Sciences Department at Imperial College London said: “Key to the success was to find a suitable detergent that yielded good protein crystals, this arduous task was facilitated greatly by a large-scale stability screen we carried out."
The ESRF and Diamond Light Source were essential to screen their crystals and collect the data used to obtain the structure. At Diamond they were also able to access specialised equipment that dehydrates the crystals, improving the resolution of their diffraction data, thus leading to much more accurate results.
“Since membrane proteins are so hard to crystallise, you have to make sure that you try everything possible to improve the quality of data you can extract from each crystal. I am very pleased that the technical effort we have put into this development has resulted in some great scientific results. We will continue to integrate this equipment to help our users with new, challenging projects.” said Dr Juan Sanchez-Weatherby, who played a key role in the development of the crystal dehydration equipment.
More information:
‘Crystal structure of a bacterial homologue of the bile acid sodium symporter ASBT’ Nien-Jen Hu, So Iwata, Alexander D. Cameron, David Drew
DOI: 10.1038/nature10450
Provided by Diamond Light Source