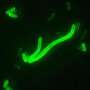
Researchers reconstruct genome of the Black Death
led by researchers at McMaster University and the University of Tubingen in Germany -- has sequenced the entire genome of the Black Death, one of the most devastating epidemics in human history.
This marks the first time scientists have been able to draft a reconstructed genome of any ancient pathogen, which will allow researchers to track changes in the pathogen's evolution and virulence over time. This work -- currently published online in the scientific journal Nature -- could lead to a better understanding of modern infectious diseases.
Geneticists Hendrik Poinar and Kirsten Bos of McMaster University and Johannes Krause and Verena Schuenemann of the University of Tubingen collaborated with Brian Golding and David Earn of McMaster University, Hernán A. Burbano and Matthias Meyer of the Max Planck Institute for Evolutionary Anthropology and Sharon DeWitte of the University of South Carolina, among others.
In a separate study published recently, the team described a novel methodological approach to pull out tiny degraded DNA fragments of the causative agent of the Black Death, and showed that a specific variant of the Yersinia pestis bacterium, was responsible for the plague that killed 50 million Europeans between 1347 and 1351.
After this success, the next major step was to attempt to "capture" and sequence the entire genome, explains Poinar, associate professor and director of the McMaster Ancient DNA Centre and an investigator with the Michael G. DeGroote Institute of Infectious Disease Research, also at McMaster University.
"The genomic data show that this bacterial strain, or variant, is the ancestor of all modern plagues we have today worldwide. Every outbreak across the globe today stems from a descendant of the medieval plague," he says. "With a better understanding of the evolution of this deadly pathogen, we are entering a new era of research into infectious disease."
"Using the same methodology, it should now be possible to study the genomes of all sorts of historic pathogens," adds Krause, one of the lead authors of the study. "This will provide us with direct insights into the evolution of human pathogens and historical pandemics."
The direct descendants of the same bubonic plague continue to exist today, killing some 2,000 people each year.
"We found that in 660 years of evolution as a human pathogen, there have been relatively few changes in the genome of the ancient organism, but those changes, however small, may or may not account for the noted increased virulence of the bug that ravaged Europe," says Poinar. "The next step is to determine why this was so deadly."
Major technical advances in DNA recovery and sequencing have dramatically expanded the scope of genetic analysis of ancient specimens, opening new horizons in the understanding of emerging and re-emerging infections.
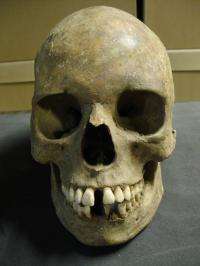
DeWitte, Bos and Schuenemann analyzed skeletal remains from victims buried in the East Smithfield "plague pits" in London, located under what is now the Royal Mint. By targeting promising specimens -- which had been pre-screened for the presence of Y. pestis -- from the dental pulp of five bodies, they were able to extract, purify and enrich specifically for the pathogen's DNA, thereby decreasing the background DNA consisting of human, fungal and other non-plague DNA.
Linking the 1349-1350 dates of the skeletal remains to the genomic data allowed the researchers to calculate the age of the ancestor of the Yersinia pestis that caused the medieval plague. This date coalesced sometime between the 12th and 13th centuries, indicating that earlier plagues such as the Justinian plague of the 6th Century -- once thought to have been caused by the same pathogen -- was likely caused by another, yet to be determined. The Justinian plague spread across the Eastern Roman Empire, killing an estimated 100 million people worldwide.
Provided by McMaster University