Rapid profiling of drug candidates
In the hunt for new medicines, any technique that expedites drug candidates into the clinic is a welcome advance. A team led by Hiroyuki Osada at the RIKEN Advanced Science Institute, Wako, recently developed a faster way to unravel the mode of action of experimental anticancer drugs, an essential step in the drug development pathway. The team—including RIKEN researchers Makoto Kawatani and Makoto Muroi—is now using this so-called ‘proteomic profiling’ technique to assess new drug leads, including a promising compound dubbed BNS-22.
Like many drug candidates, BNS-22 was developed from a natural compound. In this case, Osada and his colleagues derived BNS-22 from a compound named GUT-70, which other researchers had isolated from the Brazilian plant Calophyllum brasiliense. Since tests with GUT-70 on a culture of cancer cells revealed some activity in killing these cells, Osada and team decided to synthesize a range of closely related compounds that might exert stronger anticancer activity.
Traditionally, assessing the usefulness of novel anticancer compounds and how they work was a time-consuming process, often giving ambiguous results. Osada and his colleagues improved this process using their proteomic profiling technique, which maps protein production in cancerous cells. As cellular conditions change, including administration of an anticancer drug, so does the range of proteins that the cells produce.
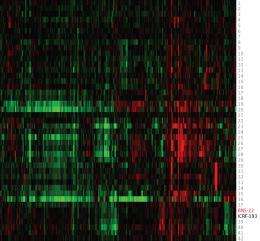
To assess BNS-22, the researchers first mapped the proteomic profiles of 42 anticancer compounds with well-established modes of action. They then compared the proteomic profile of BNS-22 to this library of profiles, using a ‘heat map’ to identify any matches (Fig. 1). Matching profiles indicate that two compounds have the same mode of action.
Osada and colleagues found that BNS-22 matches the profile of ICRF-193, a compound known to inhibit an enzyme called TOP2. This enzyme is essential for numerous cellular processes, and is a good target for anticancer drugs. Surprisingly, the findings also show that BNS-22 works in a different way from GUT-70, despite its very similar structure. “In the process of [creating derivatives of] compounds, single structural differences frequently cause a loss of activity,” Kawatani and Muroi explain. However, the team was fortunate enough to find a promising drug candidate.
To further improve the accuracy of their proteomic profiling approach, the researchers plan to further analyze their protein maps and identify significant changes in the expression of specific proteins in response to a drug. They also plan to use this technique to re-assess RIKEN’s library of natural products for new drug leads.
More information: Muroi, M., et al. Application of proteomic profiling based on 2D-DIGE for classification of compounds according to the mechanism of action. Chemistry & Biology 17, 460–470 (2010).
Kawatani, M., et al. Identification of a small-molecule inhibitor of DNA topoisomerase II by proteomic profiling. Chemistry & Biology 18, 743–751 (2011).
Provided by RIKEN