Improved method for capturing proteins holds promise for biomedical research
Antibodies are the backbone of the immune system—capable of targeting proteins associated with infection and disease. They are also vital tools for biomedical research, the development of diagnostic tests and for new therapeutic remedies.
Producing antibodies suitable for research however, has often been a difficult, costly and laborious undertaking.
Now, John Chaput and his colleagues at the Biodesign Institute at Arizona State University have developed a new way of producing antibody-like binding agents and rapidly optimizing their affinity for their target proteins. Such capture reagents are vital for revealing the subtleties of protein function, and may pave the way for improved methods of detecting and treating a broad range of diseases.
The team's results appear in today's issue of the journal ChemBioChem.
Antibodies are Y-shaped structures, capable of binding in two or more places with specific target proteins. Synthetic antibodies are much simpler forms that attempt to mimic this behavior. As Chaput explains, creating affinity reagents with strong binding properties can be accomplished by combining two weak affinity segments on a synthetic scaffold. The resulting affinity reagent, if properly constructed, can amplify the binding properties of the individual segments by two or three orders of magnitude.
"This dramatic change in affinity has the ability to transform ordinary molecules into a high affinity synthetic antibody," Chaput says. "Unfortunately, the chemistry used to make these reagents can be quite challenging and often requires a lot of trial-and-error. With NIH funding, my group has reduced the complexity of this problem to simple chemistry that is user friendly and easily amenable to high throughput automation. Such technology is absolutely necessary if we want to compete with traditional monoclonal antibody technology. "
Traditionally, antibodies for research have been extracted from animals induced to produce them in response to various protein antigens. While the technique has been invaluable to medical science, obtaining antibodies in this way is a cumbersome and costly endeavor. Instead, Chaput and his team produce synthetic antibodies that do not require cell culture, in vitro selection or the application of complex chemistry. They call their reagents DNA synbodies.
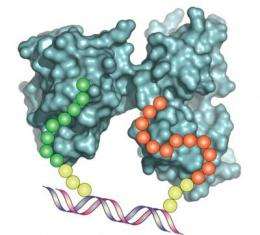
The new strategy—referred to as LINC (for Ligand Interaction by Nucleotide Conjugates) uses DNA as a programmable scaffold to determine the optimal distance needed to transform two weak affinity binding segments or ligands into a single high affinity protein capture reagent. The result is an artificial antibody, capable of binding to its antigen target with both high affinity and high specificity. The process is rapid and inexpensive. It also offers considerable flexibility, as the distance between the two ligand components bonded to the short, double-stranded DNA scaffold can be fine-tuned for optimum affinity.
In earlier work, the group identified ligand candidates by producing thousands of random sequence peptide chains—strings of amino acids, connected like pearls on a necklace. The peptide sequences were affixed to a glass microarray slide and screened against a target protein to pinpoint those that were capable of recognizing distinct protein binding sites. Two promising ligand candidates could then be combined to form a DNA synbody.
In the current study, the group instead makes use of pre-existing ligands with documented affinity for various disease-related proteins. The method involves the use of well-characterized ligands as building components for high quality DNA synbodies, eliminating the initial screening procedure and expanding the potential to tinker with the two-piece synbody in order to optimize affinity.
The peptides of choice for the study were those with high affinity for something called growth factor receptor bound protein 2 (Grb2). Grb2 has many cell-signaling functions and is an important focus of research due to its association with cellular pathways involved in tumor growth and metastasis.
By scouring the scientific literature, the group identified two peptides that recognize distinct sites on the surface of Grb2. Chaput points out, "this is a nice example where a few hours in the library can save you weeks in the lab."
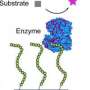
The next step was to create an assortment of synbody constructs based on these peptides. To do this, one peptide was attached to the end of a short DNA strand, while the other peptide was attached to the complementary DNA strand further along its length (see figure 1).
The two peptide strands could be attached to the scaffold in either a forward or reverse direction and could be interchanged, with either occupying the terminal end of the first DNA strand. Further, the distance between peptide segments along the DNA strands could be adjusted to yield the best target affinity.
Experiments examined binding affinity for peptide chains separated by 3, 6, 9, 12, 15 and 18 base pairs along the DNA strand, (a distance range of 1.0-6.1 nm). Inspection revealed the best results for a synbody constructed of peptides separated by 12 base pairs at a distance of 4.1 nm, compared with the other 5 constructs.
The results for the best synbody in the study were impressive, demonstrating a binding affinity five- to ten-fold stronger than commercially available antibodies for Grb2, despite the synbody's comparatively primitive architecture. In further tests, the synbody was shown to exhibit high specificity—isolating Grb2 from other proteins in a complex biological mixture and selectively binding with its target.
The technique offers a new approach to producing high qualityaffinity reagents for disease research, diagnostic testing and the development of effective therapeutics.
Provided by Arizona State University