Mussel adhesive for DNA chips
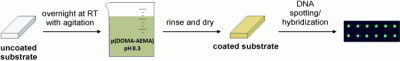
Mussels are true masters of adhesion. Whether on the wood of a pier, the metal of a ship’s hull, rocks, or to their own kind, they stick to everything. Researchers led by Philip B. Messersmith at Northwestern University (Evanston, IL/USA) have successfully synthesized a mimic of one of the "universal adhesives" used by mussels. As the scientists report in the journal Angewandte Chemie, they were able to use their synthetic "mussel glue" to fix DNA molecules on various substrates. This new, simple method seems particularly promising for the production of DNA chips for diagnostics and research.
Modern analytical strategies for the detection and analysis of biomolecules are often based on robust and inexpensive methods for immobilizing DNA, proteins, and other biomolecules on surfaces. DNA microarray techniques involve the arrangement of different DNA probes on a single chip. Various target DNA molecules are selectively fished out of the many found in a DNA sample. The target DNA is identified by means of the binding location on the chip, because the location of every probe on the chip is documented.
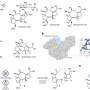
“Previous anchoring strategies have generally been developed specifically for a single substrate,” says Messersmith, “they are thus ineffective on other substrates.” Messersmith and his colleagues have now developed a universal method—inspired by mussels—that can adhere to just about any material desired. Biopolymers responsible for the unusual adhesive properties of mussels have now been identified. These polymers are rich in catechol and amino groups. “We have synthesized a catecholamine polymer that mimics the chemistry of the musselproteins,” reports Messersmith.
The new approach is rather simple: Just place the desired substrate in a solution of the catecholamine polymer overnight. The polymer adheres as a thin layer on any of the usual substrates used for DNA arrays, such as glass, as well as less common substrates such as gold, platinum, oxides, semiconductors, or various polymer substrates. The coating then easily binds DNA molecules without influencing their biological activity. This makes it possible to make micropatterns with DNA (DNA spotting), as is required for DNA chips.
The secret of the success of the catecholamine polymer: it contains special groups of atoms that can bind to a diversity of substrate materials through a variety of mechanisms. On the other hand, target DNA molecules from a sample bind exclusively to the corresponding specific DNA probes, without requiring a treatment to block unspecific binding to the substrate. Says Messersmith: “The new coating strategy may significantly simplify DNA microarray technology.”
More information: Phillip B. Messersmith, Facile DNA Immobilization on Surfaces through Catecholamine Polymer, Angewandte Chemie International Edition, dx.doi.org/10.1002/anie.201005001
Provided by Wiley