This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers uncover natural variation in wild emmer wheat for broad-spectrum disease resistance
Bread wheat is one of the most important staple crops for millions of people and is apparently the largest cultivated and traded cereal worldwide. Bread wheat is a hexaploid species with three subgenomes (2n = 6x = 42, AABBDD) that has undergone two separate allopolyploidization and domestication events.
Due to the bottleneck effects, modern wheat suffers from extremely low genetic diversity, which has led to genetic erosion and increased susceptibility and vulnerability to environmental stresses, pests, and diseases. Wild emmer wheat, Triticum dicoccoides, (2n = 4x = 28, AABB), is the direct wild ancestor of both durum and bread wheat, providing novel natural variation for modern wheat improvement.
Researchers from the Institute of Genetics and Developmental Biology (IGDB) of the Chinese Academy of Sciences have made progress in cloning a broad-spectrum powdery mildew resistance gene, Pm36, encoding a novel tandem kinase with a transmembrane domain (WTK7-TM) originating from wild emmer wheat.
The results were published online in Nature Communications on April 10.
Pm36 was first identified from wild emmer-durum wheat backcross introgression lines and mapped to the 5BL chromosome arm by scientists of University of Bari, Italy, using amplified fragment length polymorphism, simple sequence repeat (SSR), and expressed sequence tag (EST)-derived makers.
Scientists from China Agricultural University developed powdery mildew resistance wild emmer-common wheat backcross introgression line in about the same period of time and fine mapped the same gene using polymorphic SSR, EST-derived sequence tagged site markers by performing comparative genomic analysis.
In this study, to clone Pm36, a larger segregation population consisting of 31,786 gametes was used to fine map Pm36 in a small genomic interval containing only four high-confidence genes according to the available reference genome sequences of wheat and its wild relatives. However, none of these genes was found to be Pm36 after the two expressed genes within this interval were separately introduced into the highly susceptible wheat cultivar Fielder by Agrobacterium-mediated transformation for functional characterization.
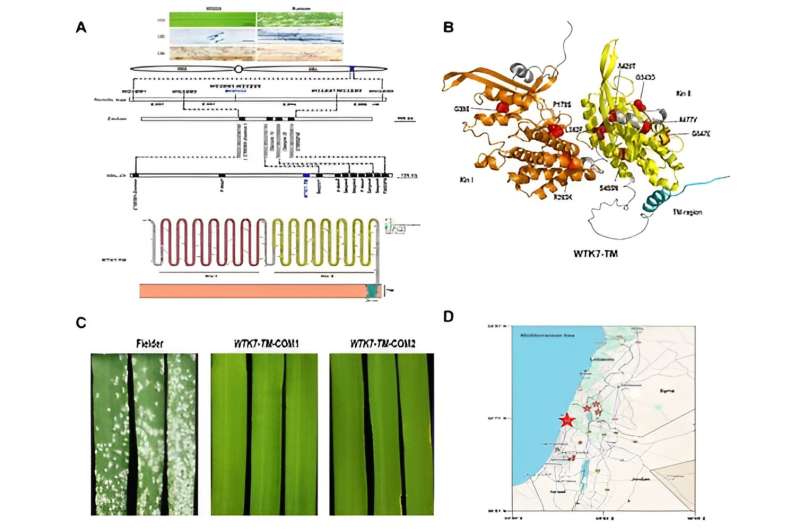
The researchers then wondered whether this was a variation in the genomic structure of the Pm36 gene region. Taking advantage of the reduced cost of PacBio SMRT long-read sequencing technology, the tetraploid wild emmer-durum introgression line 5BIL-29 carrying Pm36 was sequenced to generate HiFi reads and Hi-C reads for genome sequence assembly. A 7.1 Mb contig ptg000422 spanning the entire Pm36 physical mapping interval (1.17 Mb) was captured from the assembled genome.
Not surprisingly, a large sequence insertion was found in the Pm36 physical region harboring seven additional predicted genes and a putative tandem kinase with a predicted transmembrane domain (WTK7-TM) was further confirmed as the Pm36 gene.
Haplotypic variation analysis revealed that Pm36 (WTK7-TM) was presented only in the southern wild emmer gene pool. The absence of Pm36 and several other cloned disease resistance genes, such as Pm41, Yr15 and Yr36, in wild emmer wheat natural habitats from southeastern Turkey, where wheat is believed to have been domesticated, as well as in cultivated tetraploid and hexaploid wheat, revealed that these disease resistance genes were left behind in the wild and not integrated into the cultivated wheat gene pool.
Most of the cloned wheat resistance genes encode for intracellular nucleotide-binding leucine-rich repeat receptor proteins. Wheat tandem kinase (WTK) was recently characterized as a novel resistance protein in wheat and wild relatives for many types of disease resistance, including stripe rust, stem rust, leaf rust, powdery mildew, and wheat blast.
Pm36/WTK7-TM was discovered in wild emmer wheat and has been shown to be resistant to diverse isolate of Blumeria graminis f. sp. tritici. Pm36 has been used to develop advanced breeding lines with broad spectrum resistance and high yield potential to ensure food security.
More information: Miaomiao Li et al, A membrane associated tandem kinase from wild emmer wheat confers broad-spectrum resistance to powdery mildew, Nature Communications (2024). DOI: 10.1038/s41467-024-47497-w
Journal information: Nature Communications
Provided by Chinese Academy of Sciences