This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Scientists reveal how SID-1 recognizes dsRNA and initiates systemic RNA interference
RNA interference (RNAi) is a fascinating biological process in worms, plants, fungi, and metazoans that has been a valuable tool for studying gene function and as therapeutics.
In Caenorhabditis elegans, the multipass transmembrane protein, systemic RNA interference defective protein 1 (SID-1), plays an indispensable role in the uptake and delivery of double-stranded RNA (dsRNA) between cells and tissues, leading to systemic RNAi.
In addition, two human SID-1 homologs, SID1 transmembrane family member 1 (SIDT1) and SIDT2, have been implicated in RNA transport. However, the underlying molecular mechanisms of how SID-1 specifically distinguishes dsRNA from single-stranded RNA (ssRNA) and DNA and facilitates subsequent dsRNA transport between cells remain unknown.
Answers to these questions are important for understanding systemic RNAi and for aiding in RNA-related applications.
Dr. Zhang Jiangtao in Prof. Jiang Daohua's group from the Institute of Physics of the Chinese Academy of Sciences, has demonstrated how SID-1 specifically recognizes dsRNA and provided important insights into the internalization of dsRNA by SID-1 by combining cryo-EM, in vitro, and in vivo experiments. The work is published in the journal Nature Structural & Molecular Biology.
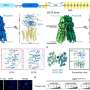
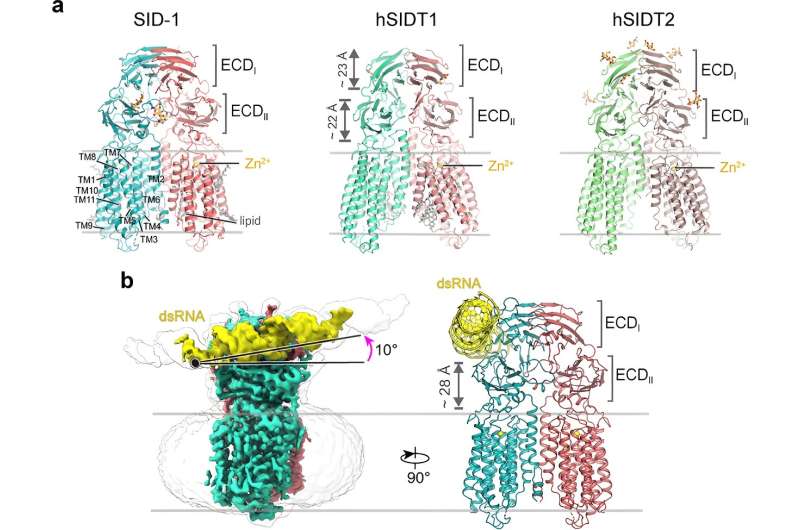
For more than two decades, SID-1 was thought to function as a dsRNA channel. Here, the researchers solved high-resolution cryo-EM structures of SID-1 and the human SID-1 homologs SIDT1 and SIDT2, revealing the conserved architecture of C. elegans and human SID-1 homologs.
The SID-1 homologs are organized in a homo-dimeric fashion. Surprisingly, the SID-1 dimer does not show an obvious pore within the transmembrane domain, suggesting that SID-1 may not function as a dsRNA channel. MST binding assays confirmed that SID-1 can potently and specifically bind to dsRNA but not to dsDNA.
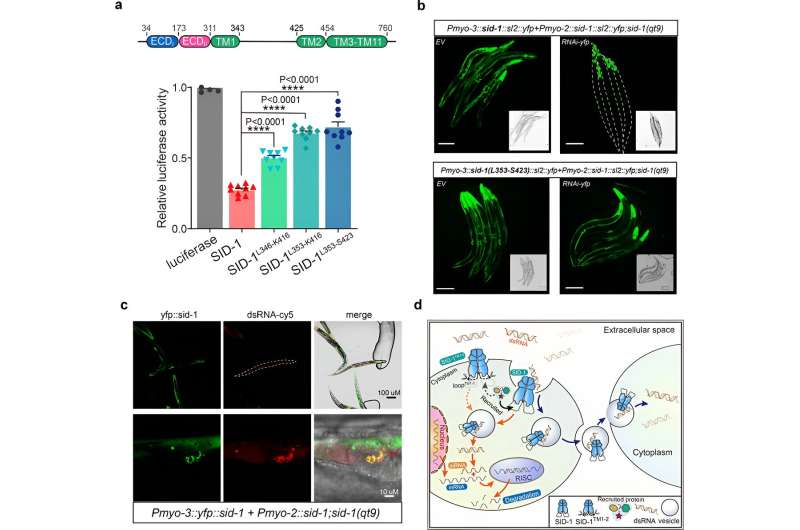
Subsequently, the researchers obtained the cryo-EM structure of the SID-1-dsRNA complex, demonstrating the detailed dsRNA binding mode and the molecular determinants of how SID-1 distinguishes dsRNA from ssRNA and DNA.
Interestingly, such determinants are not present in human SIDT1 or SIDT2. The structural findings were supported by mutagenesis studies using MST binding assays, dsRNA uptake in S2 cells, and in vivo systemic RNAi assays.
Finally, the researchers show that the removal of the long intracellular loop transmembrane helices 1 and 2 did not affect SID-1 dimerization, cell localization, or dsRNA binding, but it significantly impaired dsRNA uptake in S2 cells and systemic RNAi in C. elegans.
Furthermore, co-localization revealed that SID-1 and dsRNA co-locate in vesicle-like subcellular organelles. Based on these results, the researchers propose that SID-1 functions as a dsRNA receptor and facilitates subsequent dsRNA internalization by recruiting endocytic-related proteins via the long loop.
More information: Jiangtao Zhang et al, Structural insights into double-stranded RNA recognition and transport by SID-1, Nature Structural & Molecular Biology (2024). DOI: 10.1038/s41594-024-01276-9
Journal information: Nature Structural & Molecular Biology
Provided by Chinese Academy of Sciences