This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
A powerful technique for tracking a protein's fleeting shape changes
Researchers at Weill Cornell Medicine have developed a powerful, new technique to generate "movies" of changing protein structures and speeds of up to 50 frames per second.
Senior author, Dr. Simon Scheuring, the Distinguished Professor of Anesthesiology Research at Weill Cornell Medicine and colleagues developed the new approach to gain a better understanding of how biological molecules change structurally over time.
Although investigators in this field routinely image static proteins and other molecules finely enough to resolve the positions of individual atoms, the resulting structural pictures or models are snapshots. Recording the dynamics of molecular structures—making movies—has been a much harder challenge. The lead author of the study is Yining Jiang, a doctoral candidate in the Weill Cornell Graduate School of Biomedical Sciences.
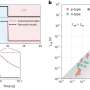
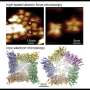
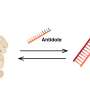
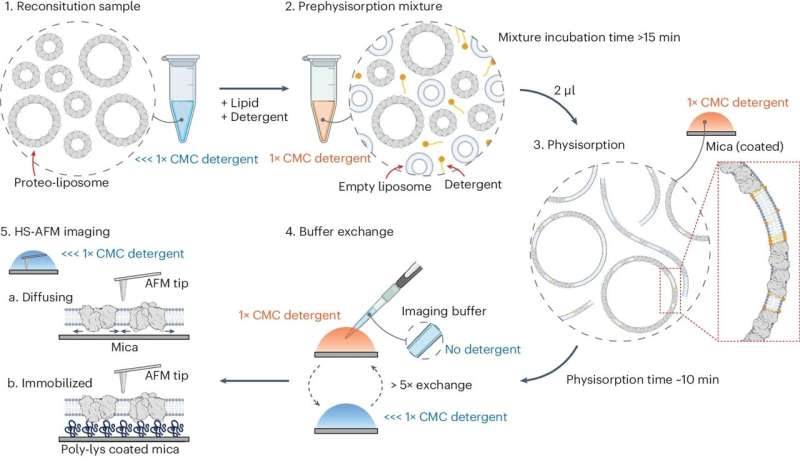
In their study, published April 17 in Nature Structural & Molecular Biology, the researchers used a relatively new measurement technique called high-speed atomic-force microscopy (HS-AFM), which employs an extremely sensitive probe to scan across molecules' surfaces, essentially feeling their structures. As a key innovation, the scientists found a method to isolate their target molecule, a single protein, thus avoiding effects from protein-to-protein interactions and enabling faster and more precise scanning.
The researchers applied their new single-molecule HS-AFM approach to a protein called GltPh, a "transporter" that sits in the cell membrane, directing neurotransmitter molecules into the cell. Such transporters are among the favorite targets of structural biologists due to their complex and puzzling dynamics, and their importance in health and disease.
The researchers obtained dynamic structural data on GltPh with an unprecedented combination of high spatial and time resolution—and stability, so that they could record tiny fluctuations in GltPh's structure continuously for minutes.
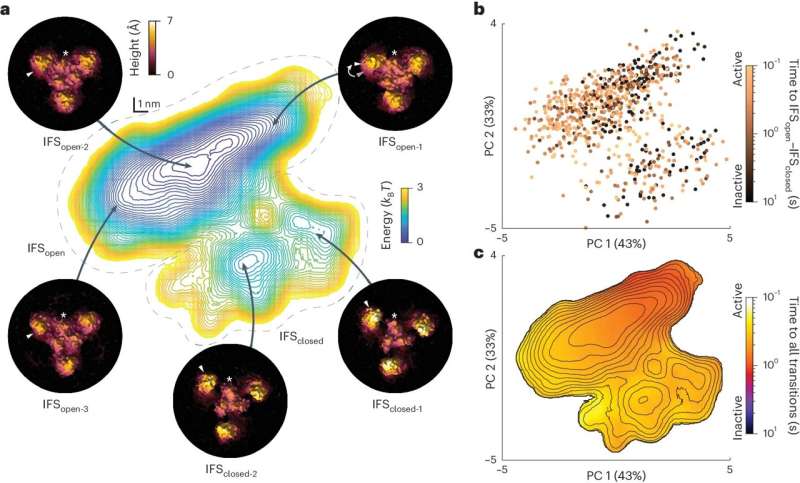
An unsolved phenomenon in such proteins was termed "wanderlust" kinetics, meaning that molecules were reported to functionally change between high and low activity modes, for no obvious reason. The work revealed a previously unseen structural state of GltPh, in which the transporter is locked and functionally asleep, uncovering the basis of 'wanderlust' kinetics.
The researchers emphasized that their new approach, which they are continually trying to optimize, is generalizable for studying other proteins, including membrane-embedded proteins. Overall, they said, this work opens up new possibilities to track the precise structure of a protein moment-by-moment during its cycles of activity and rest.
More information: Yining Jiang et al, HS-AFM single-molecule structural biology uncovers basis of transporter wanderlust kinetics, Nature Structural & Molecular Biology (2024). DOI: 10.1038/s41594-024-01260-3
Journal information: Nature Structural & Molecular Biology
Provided by Weill Cornell Medical College