This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
RNA splicing regulation discovery provides insight into bone diseases
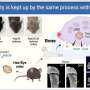
In today's aging societies, bone and joint diseases are becoming increasingly common. For example, in Japan alone, over 12 million people suffer from osteoporosis, a condition that severely weakens bones and makes them fragile. If we are to find effective treatments for such disorders, understanding the cellular processes involved in the maintenance of bone and joint tissue is an essential first step.
Osteoclasts are a particularly important type of cell involved in bone maintenance. These cells absorb old or damaged bone and digest it, allowing the body to reuse necessary materials like calcium and giving way to new bones. As one might expect, various bone diseases arise when osteoclasts do not fulfill their role properly. Scientists have been investigating the mechanisms that regulate the proliferation and differentiation of precursor cells into osteoclasts.
Interestingly, in a study published in 2020, researchers from Tokyo University of Science (TUS) led by Professor Tadayoshi Hayata revealed that the cytoplasmic polyadenylation element-binding protein 4 (Cpeb4) protein is essential in osteoclast differentiation.
They also discovered that this protein, which regulates the stability and translation of messenger RNA (mRNA) molecules, was transported into specific structures within the cell's nucleus when osteoclast differentiation was induced. However, just how this relocation occurs and what Cpeb4 exactly does within these nuclear structures still remains a mystery.
Now, in a recent study published in the Journal of Cellular Physiology on 29 January 2024, Prof. Hayata and Mr. Yasuhiro Arasaki from TUS tackled these knowledge gaps. Motivated by the intricate and complex process of osteoclast differentiation, they sought to more thoroughly understand how the "life cycle" of mRNA, i.e., mRNA metabolism, is involved.
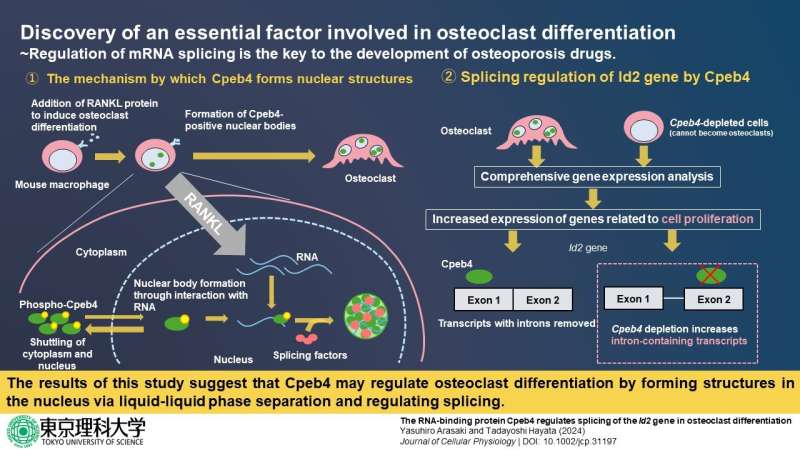
First, the researchers introduced strategic modifications to Cpeb4 proteins and performed a series of experiments in cell cultures. They found that the localization of Cpbe4 in the abovementioned nuclear bodies occurred owing to its ability to bind to RNA molecules.
Afterward, seeking to understand the role of Cpeb4 in the nucleus, the researchers demonstrated that Cpeb4 co-localized with certain mRNA splicing factors. These proteins are involved in the process of mRNA splicing, which is a key step in mRNA metabolism. Put simply, it enables a cell to produce diverse mature mRNA molecules (and eventually proteins) from a single gene.
Through RNA sequencing and gene analysis in Cpeb4-depleted cells, they found that Cpeb4 alters the expression of multiple genes associated with splicing events in freshly differentiated osteoclasts. Finally, further experiments revealed that Cpeb4 only altered the splicing patterns of Id2 mRNA, an essential protein known to regulate osteoclast differentiation and development.
Overall, this study sheds important light on the mechanisms that regulate osteoclast differentiation.
"Through this research, we were able to identify important factors involved in regulating mRNA splicing during the osteoclast differentiation process and obtained new knowledge regarding the control of mRNA splicing during osteoclast differentiation," comments Prof. Hayata.
While the contribution of Cpeb4 is smaller than that of RANKL, a signaling factor that induces osteoclast differentiation, targeting Cpeb4 may have the advantage of reducing the side effects of existing drugs as too much inhibition of osteoclast differentiation with RANKL inhibitory antibodies would halt bone remodeling.
Importantly, the results contribute to a more detailed understanding of how bones are maintained. "Although we used cultured mouse cells in our study, there are also research reports that show a correlation between variations in the CPEB4 gene and bone density in humans," says Prof. Hayata. "We hope that our findings will help clarify the relationship between these two in the near future."
Most importantly, the present study's findings may prove to be a crucial stepping stone for advancing diagnostic techniques and treatments for bone and joint diseases. GWAS analysis has reported a correlation between single nucleotide polymorphisms in introns of the CPEB4 gene region and the estimated bone density. Therefore, it is possible that CPEB4 expression and activity can be used as diagnostic criteria.
However, the researchers note that it is unclear whether Cpeb4 actually regulates bone metabolism in vivo. Therefore, clarification of the molecular basis of Cpeb4 in bone metabolism in mice would help to establish a therapeutic approach. Additionally, recent studies have reported that Cpeb4 is expressed in various cancer cells and contributes to cancer cell survival. In cancer, Cpeb4 contributes to mRNA stability, although splicing regulation may exist.
"The discovery of part of the mechanisms by which Cpeb4 controls osteoclast differentiation could lead to the elucidation of pathologies, including osteoporosis and rheumatoid arthritis, and ultimately become the foundation for the development of new therapeutic drugs," concludes Prof. Hayata.
More information: Yasuhiro Arasaki et al, The RNA‐binding protein Cpeb4 regulates splicing of the Id2 gene in osteoclast differentiation, Journal of Cellular Physiology (2024). DOI: 10.1002/jcp.31197
Provided by Tokyo University of Science