This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
RNA-dependent protein research advances the fight against malaria
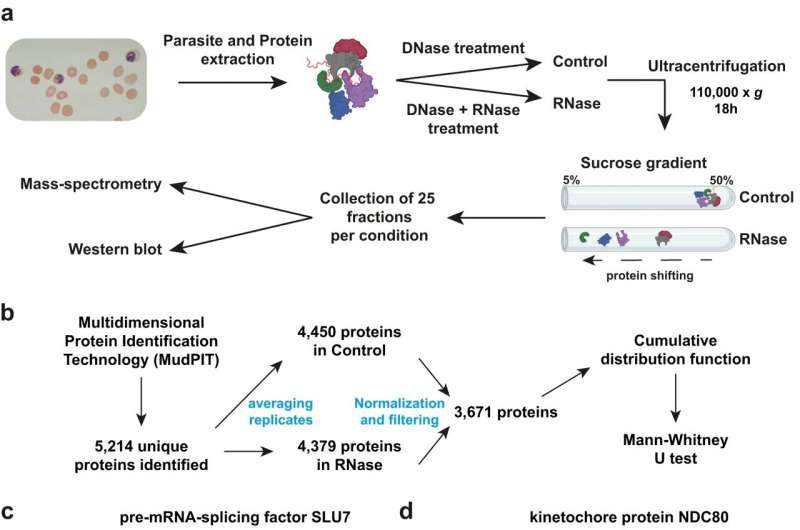
New work by a team led by scientists at the University of California, Riverside, has taken research one step closer to designing new therapies to fight and eradicate malaria thanks to a lab technique called R-DeeP.
The team is studying RNA-dependent proteins—assemblies of RNA molecules and proteins that are critical for cell survival. These RNA-protein complexes play fundamental roles in many cellular processes.
In Plasmodium falciparum, the deadliest human malaria parasite, however, scientists have been able to identify and characterize only a limited number of RNA-dependent proteins due to the complexity of the parasite life cycle progression and limited tools available to edit the parasite genome efficiently.
Now, using a comprehensive "molecular search tool" called R-DeeP, Karine Le Roch, and her colleagues have identified and characterized 898 RNA-dependent proteins in P. falciparum, including uncharacterized and parasite-specific proteins, which could lead to novel therapeutic targets against malaria. Study results appear in Nature Communications.
"We also validated that one novel parasite-specific RNA-binding protein, PF3D7_0823200, interacts with various Plasmodium transcripts involved in controlling virulence," said Le Roch, a professor of molecular, cell and systems biology and director of the UCR Center for Infectious Disease and Vector Research. "This RNA-binding protein could be targeted by new drugs and is, therefore, of interest in the fight against malaria."
During transcription, a gene's DNA sequence is copied by enzymes to make an RNA molecule. Of the 898 RNA-dependent proteins the researchers identified, only 39% of them had already been identified as associated with RNA.
"Our study provides the first snapshot of the Plasmodium protein-protein and protein-RNA interaction network in the parasite," Le Roch said. "These generated R-DeeP results highlight the importance of RNA in many biological pathways in the parasite and identify new targets for combating malaria."
More information: Thomas Hollin et al, Proteome-Wide Identification of RNA-dependent proteins and an emerging role for RNAs in Plasmodium falciparum protein complexes, Nature Communications (2024). DOI: 10.1038/s41467-024-45519-1
Provided by University of California - Riverside