This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
New research brings order to disordered proteins
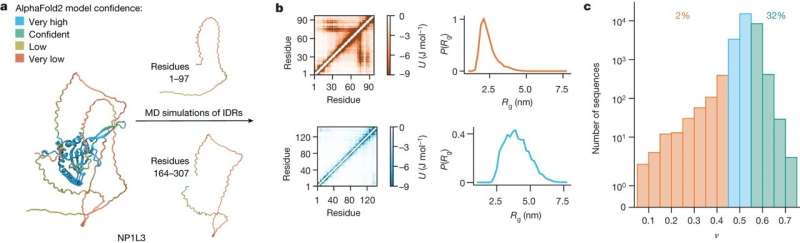
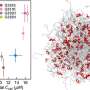
Protein molecules lie at the heart of biology. Our typical understanding of proteins states that each type of protein has a specific three-dimensional shape that enables it to perform its function. This dogma is challenged by intrinsically disordered proteins, which make up one-third of all proteins and have central biological functions even though their shapes are constantly changing.
Until now, our understanding of the structural properties of this intriguing class of proteins has been based on studies of only a small number of examples. In research published today in the journal Nature, researchers from the Department of Biology, University of Copenhagen have shown how all (approximately 28,000) disordered proteins in the human body behave.
"I have always been fascinated by intrinsically disordered proteins because they seem to defy most of the rules of how a protein should behave. For the last 20 years, we have worked on figuring out how these strange proteins look and whether new rules need to be applied to describe them. For the first time, we have now been able to study the structure of all human disordered proteins and begun to provide order into this world of molecular disorder," says Professor Kresten Lindorff-Larsen, director of the NNF center PRISM, in which the research was performed.
The goal of the PRISM center is to combine computational methods from biophysics and machine learning, with methods from cell biology to study how genetic variants cause disease. But until now, the researchers did not know how most disordered proteins looked, and hence could not even begin to study how mutations in the genes encoding for them might be able to cause disease.
Until recently, we examined the disordered proteins one-by-one, and it was essential to find a way to study them on a larger scale," says Assistant Professor Giulio Tesei, who is one of the lead authors of the new paper. "We came up with an approach where we could use experimental measurements on disordered proteins to develop a computational model to predict their properties. Since this model is both accurate and fast, we can now look at them all."
The study was co-led by bachelor student, Anna Ida Trolle, who says, "When I started the project, I didn't know that you typically just study one or two proteins at a time. So, when Giulio and Kresten suggested that I should study some 28,000 proteins, I fortunately didn't realize how crazy an idea it was. However, we quickly found a way to generate and keep track of the large amount of data and were able to use it to study the biology and evolution of disordered proteins.
Lindorff-Larsen concludes, "This has been a challenging but also an extremely fun project, that was only made possible by the contributions of several people with diverse expertise in the PRISM center. We have made new steps on linking the molecular properties of disordered proteins to their biological function and roles in disease. Finally, we are beginning to understand the language of disordered proteins."
More information: Giulio Tesei et al, Conformational ensembles of the human intrinsically disordered proteome, Nature (2024). DOI: 10.1038/s41586-023-07004-5
Provided by University of Copenhagen