This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
proofread
The metabolism of bacteria: New method reveals host-microbe interactions
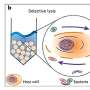
The fascinating world of bacteria that live as symbionts or parasites in animal hosts often remains a mystery to researchers. Under the leadership of Prof. Dr. Manuel Liebeke, Kiel University (CAU) and the Max Planck Institute for Marine Microbiology in Bremen are contributing to solving this puzzle by researching the interactions between microbes and their host. However, there has been a lack of insight into what bacteria do in their natural environment.
Often, bacteria cannot be cultured in the laboratory, and researchers must rely on information from the bacterial genome obtained from environmental samples to gain theoretical insights into the metabolism of microorganisms. However, there has been a lack of insight into what they do in their natural environment. To solve this puzzle, scientists began researching the metabolome of bacteria—everything that has to do with their metabolism, including metabolites such as sugars or fats.
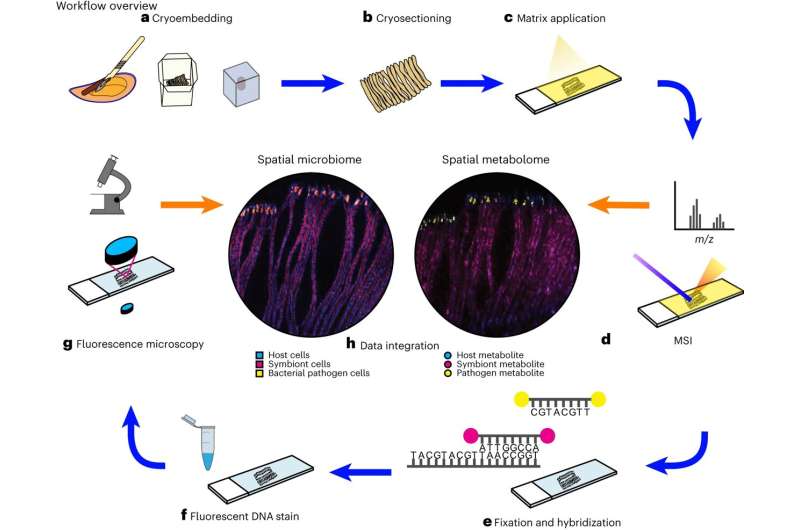
In a pioneering study, Liebeke's team developed a method to identify individual bacteria and simultaneously determine which metabolites are present in the cells without cultivating the bacteria in the laboratory. This method allows them to study how bacteria live and survive as symbiotic subtenants, such as mussels. The team analyzed hundreds of metabolites on an area smaller than one square millimeter. In September, the Kiel and Bremen researchers published their results in Nature Protocols.
A frozen moment enables detailed observation
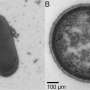
"We create a snapshot, so to speak, of the bacteria at work, exactly as they are active in their natural environment, particularly within an animal cell," Liebeke explained. "And we can do that at an impressive resolution of a few micrometers, about ten times thinner than a human hair."
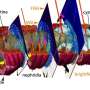
A special feature of this method is the use of flash-frozen tissue, which is cut wafer-thin. The researchers then use a special mass spectrometry technique called MALDI-MS imaging to create a snapshot of the chemical compounds in the cells.
However, drawing the correct conclusions from the images of the metabolites is only possible if they know which bacteria produce or use them. To solve this problem, the researchers also use fluorescence in situ hybridization (FISH) to identify and localize individual bacterial cells in the sample.
"Applying this method to host-microbe communities will give us many exciting new insights into chemical communication between organisms," said Patric Bourceau from the Max Planck Institute for Marine Microbiology, lead author of the protocol developed to apply the method.
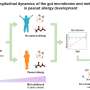
This work opens new doors for studying bacteria and their interactions with their host. In addition, the method presented here also offers promising potential applications for the future: developed at the Max Planck Institute in Bremen, Liebeke's new working group at the CAU is now using it to study the human gut microbiome and its influence on metabolism. For example, this could help us better understand inflammatory bowel diseases. With the publication of a detailed protocol, the application of the technique is now open to other researchers worldwide.
In summary, applying microscopy and metabolomics (the research field dedicated to studying metabolites) provides insights into host-microbe interactions' functional and chemical ecology. The steady advances in MALDI-MSI technology make it possible to illustrate microbial colonies, biofilms, individual eukaryotic cells, and even bacterial microcolonies. Today, MALDI-MSI technology is on the verge of being able to provide images of individual bacterial cells. The protocol presented here forms the basis for analyzing and understanding metabolic interactions down to the micrometer.
More information: Patric Bourceau et al, Visualization of metabolites and microbes at high spatial resolution using MALDI mass spectrometry imaging and in situ fluorescence labeling, Nature Protocols (2023). DOI: 10.1038/s41596-023-00864-1
Journal information: Nature Protocols
Provided by Christian-Albrechts-Universität zu Kiel