This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Potent anti-cancer therapy created using 'click chemistry'
A potent anti-cancer therapy has been created using Nobel prize-winning "click chemistry," where molecules click together like LEGO bricks, in a new study by UCL and Stanford University researchers.
The study, published in Nature Chemistry, opens up new possibilities for how cutting-edge cancer immunotherapies might be built in future.
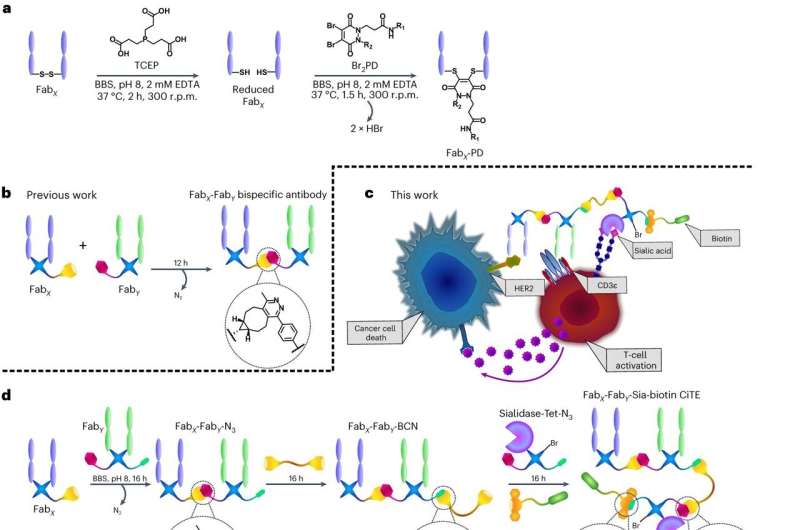
The research team created an anti-cancer therapy with three components: one targeting the cancer cell, another recruiting a white blood cell called a T cell to attack the cancer cell, and a third knocking out part of the cancer cell's defenses.
Previously, this type of three-component therapy has only been built using a complex process called protein engineering, in which DNA sequences for multiple proteins are combined and inserted into a single cell.
One of the three-component therapies the researchers built, which used an enzyme called sialidase to strip away sugars that the cancer cell uses to hide itself, was especially effective at killing breast cancer cells in a dish. The researchers said this showed that the enzyme—which only recently started being explored in cancer research—has the potential to be the basis of next-generation anti-cancer agents.
First author Dr. Peter Szijj (UCL Chemistry) said, "Click chemistry is a quicker and more adaptable way to build these multifunctional anti-cancer agents than protein engineering. It's relatively easy to attach click 'handles' to proteins so you can try lots of combinations quickly to test what might work best. Using protein engineering, you need a separate mechanism for each component."
Senior corresponding author Professor Vijay Chudasama (UCL Chemistry) said, "As proteins are large and complex molecules, you require a combination of precise protein modification and reliable click chemistry to attach them together in a controlled manner. We have achieved this and shown our strategy to be an interesting alternative to using the classical protein engineering approach."
"We hope that by using chemistry to create novel and highly sophisticated multi-protein anti-cancer agents we can inspire chemists to cross the typical boundaries of the discipline to engage in novel applications in areas such as medical imaging, diagnostics and disease therapies."
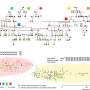
Click chemistry relies on two reaction partners (click handles) that can attach to each other very rapidly and selectively, without the production of any toxic by-products. These click handles can be added to proteins, in this case using functionalized pyridazinediones (PDs), allowing the proteins to click neatly together like LEGO.
The pioneers of click chemistry were awarded the 2022 Nobel Prize in Chemistry. Professor Carolyn R. Bertozzi, of the University of Stanford, who is a co-author on this latest paper, was one of three winners of the prize for her work on biorthogonal chemistry—click chemistry in living cells.
For the new paper, researchers at UCL first clicked two antibody fragments together—one fragment binding to a cancer cell, another fragment binding to a T cell so that it would destroy the cancer cell. Similar T cell engagers, created via protein engineering, have already been approved for use in humans and are used to treat cancers such as multiple myeloma, a rare blood cancer, in the United States and Europe.
The team then added a third component, a checkpoint inhibitor, which removes an aspect of a cancer cell's defenses. This component was either a PD-1-blocking antibody fragment, which is already used to treat specific advanced forms of skin or lung cancer and re-awakens immune cells to target cancer cells; or the more experimental sialidase enzyme, which strips away specific sugars (sialic acids) on the surface of the cancer cell as well as on the T cell.
These sugars, present on all our cells, are produced in large amounts by cancer cells and help them to hide from our immune system by switching off approaching immune cells.
The research team found that adding either of these components improved the cancer-killing efficiency of the therapy, and that adding sialidase was especially potent.
The researchers also added a fourth molecule, biotin, allowing them to visualize how well the components bound to their respective targets. They said that this could be substituted for another small molecule with a different function—for instance, to minimize side-effects by masking the protein construct until it reaches its intended target: the cancer.
In the paper, the researchers said that using chemistry in this way to create cancer therapies showed "much untapped potential that is still waiting to be uncovered."
This sialidase enzyme-containing therapeutic now needs to be tested in animals before any trials involving humans could begin.
More information: Peter A. Szijj et al, Chemical generation of checkpoint inhibitory T cell engagers for the treatment of cancer, Nature Chemistry (2023). DOI: 10.1038/s41557-023-01280-4
Journal information: Nature Chemistry
Provided by University College London