This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers invent trap for capturing and comparing individual bacterial cells
All hospitals battle an invisible threat: Pseudomonas aeruginosa. It is a type of bacteria that affects thousands of patients each year in intensive care units, where it can cause sepsis, pneumonia and other types of infections.
"For the average healthy person, P. aeruginosa does not pose a serious threat," said University of Notre Dame bacteriologist Joshua Shrout. "But for those who are most vulnerable—who are immunocompromised, who are using a ventilator or catheter, or who are recovering from serious burns or surgeries—it is not just serious, but life-threatening. And that is due to the bacteria's sophisticated suite of self-defense tactics."
Shrout, a professor in the Department of Civil and Environmental Engineering and Earth Sciences, leads a research group that studies those tactics in intricate detail. He explains that in addition to being resistant to many of the most common antibiotics, P. aeruginosa readily sticks to surfaces where it creates its own protection by blanketing itself in a polymer-like biofilm. Under certain conditions, P. aeruginosa can also acquire other organisms' antibiotic strains, and it can even generate cyanide to kill off competitors.
Thinking outside the Petri dish
A decade ago, Shrout began collaborating with Paul Bohn, the Arthur J. Schmitt Professor of Chemical and Biomolecular Engineering and director of Notre Dame's Berthiaume Institute for Precision Health. Bohn's lab specializes in creating new technologies for more precise analysis of cells and molecules. Together, Bohn and Shrout began searching for new ways to observe microorganisms like P. aeruginosa, moving beyond the traditional process of observing cell cultures grown in a Petri dish.
"If you grow and observe a whole culture of cells, you are seeing general behaviors, on average," Bohn said. "But we now know that the biggest effects sometimes come from the minority of a given population."
Shrout explained this effect using an analogy with other organisms' behaviors. "Imagine you are sitting on your back porch listening to the crickets outside," he said. "There might be thousands of crickets, but the ones that affect you are the ones chirping. There can be similar types of variation in populations of bacteria, too. However, most microbiologists don't have the tools to fully appreciate the effects of minor players within a population."
Bohn and Shrout knew that before they could find the key differences in a population of P. aeruginosa, they would need to solve some major technical challenges. First, they would need to capture individual cells. Then, to conduct a meaningful experiment, they would need to apply the same stimulus to the cells in the same way at the same time. And, finally, they would need a way to observe the experiment, tracking the differences between the cells as they respond to the stimulus.
With funding from the National Institute of Allergy and Infectious Diseases and the National Science Foundation, Bohn and Shrout decided to pioneer a new approach. "At the start," Bohn said, "we were captivated by a very simple question: 'What if we drill holes the same diameter of the bacterial cells? If we do, could we induce the cells to move in and stay inside?'"
Building a better bacteria trap
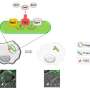
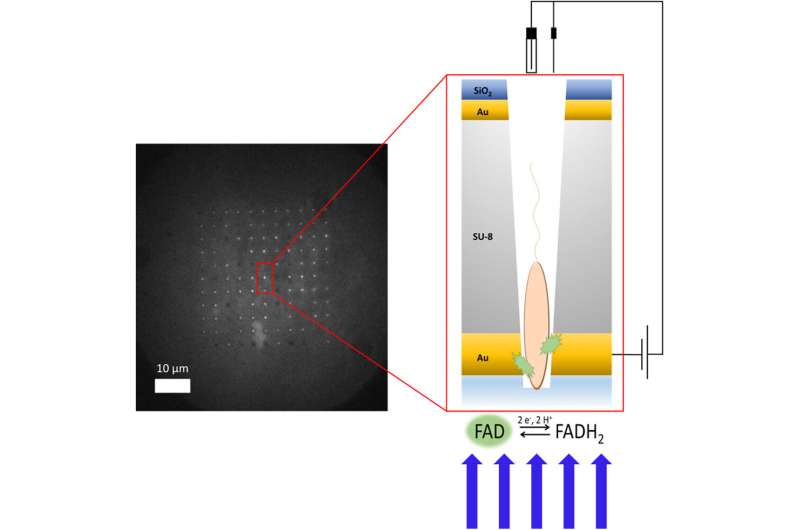
Allison Cutri, a chemistry doctoral student in Bohn's research group, took on the challenge of creating the device as part of her dissertation research. Cutri worked alongside postdoctoral research associate Vignesh Sundaresan to develop the main platform of the device inside the clean room at Notre Dame's Nanofabrication Facility. They used an additive process, building the platform layer by layer like a microchip. For the core of the device, they used a layer of epoxy, and they capped both sides with a thin veneer of gold to conduct electricity to targeted points on each bacterium, like attaching an electrode at either end of the cell.
Then Cutri worked with Notre Dame's Integrated Imaging Facility to mill more than 100 tiny holes called micropores into the device. Each micropore is drilled using a focused ion beam to a diameter of less than a micron—or about 50 times smaller than the diameter of a human hair—to allow one bacterium to slip inside.
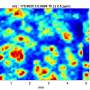
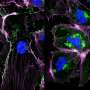
Thanks to P. aeruginosa's proclivity to stick to surfaces, Cutri was able to corral individual cells onto the device and into the micropores, where each settled into its own pore. The team was then able to apply electrical charges through the layers of gold while filming the experiment through a fluorescence microscope specially designed to look at one class of bacterial molecules.
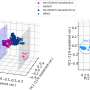
They observed glowing from the wells, which proved the device had been properly loaded with bacteria. And as they tested regiment after regiment of cells, they noticed that patterns began to emerge. Some cells glowed along with the electrical charge. Others glowed intermittently in response to the charge, while a third set glowed independently of the charge.
"We are still not sure what other differences might exist or what the differences mean for combatting P. aeruginosa," Shrout said. Upon further investigation, however, they were able to see similar patterns in a different type of bacteria: E. coli. They were also able to determine that each cell's metabolic state—whether it was taking in or expending energy—played a key role in shaping how it behaved.
For now, the researchers said they are hopeful that the patterns they observed and the device they created, which were recently described in Cell Reports Physical Science, will inspire similar research projects.
"We have known for some time that P. aeruginosa exhibits fluorescent behavior in response to a charge," Cutri explained. "But to recognize that there is so much variation in the ways that cells respond, you need a device that will allow you to observe cells individually. With existing approaches, those differences would be masked."
Shrout added, "As a researcher, it is rewarding to not just be asking new questions but to also be developing the new tools and new platforms that make those questions possible. By bringing a knowledge of cell behavior together with nanofabrication and high-resolution imaging, that is what we have been able to do with this project."
More information: Allison R. Cutri et al, Spectroelectrochemical behavior of parallel arrays of single vertically oriented Pseudomonas aeruginosa cells, Cell Reports Physical Science (2023). DOI: 10.1016/j.xcrp.2023.101368
Journal information: Cell Reports Physical Science
Provided by University of Notre Dame