This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Innovative approach opens the door to COVID nanobody therapies
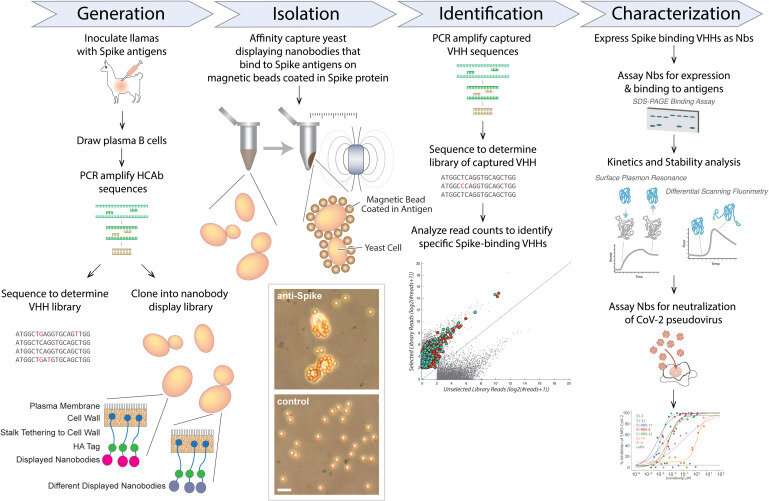
COVID is not yet under control. Despite a bevy of vaccines, monoclonal antibodies, and antivirals, the virus continues to mutate and elude us. One solution that scientists have been exploring since the early days of the pandemic may come in the form of tiny antibodies derived from llamas, which target various parts of the SARS-CoV-2 spike protein.
In a new study in the Journal of Biological Chemistry, researchers describe a less expensive way to isolate and identify these so-called nanobodies. The findings will make it easier for scientists around the world to try their hand at discovering nanobodies that target SARS-CoV-2 or other viruses.
"Our method is more straightforward and less expensive than existing techniques," says Rockefeller's Michael P. Rout. "You do need a llama, but that—along with all the most complicated parts of the process—can be outsourced."
The authors have already used this optimized method to identify multiple nanobodies that appear to work against key variants of the virus, including omicron. "COVID is clearly going to be a problem for some time," Rout says. "We show that many of the nanobodies we have identified with this method target variants-of-concern, so they have real therapeutic potential."
Nanobody novelty
Nanobodies may work where larger antibodies fail, in part due to their compact size. Studies have shown that nanobodies can squeeze into parts of the SARS-CoV-2 virus that larger antibodies cannot reach. Nanobodies also have unusually long shelf-lives, cost very little to mass-produce, and because of their unique physical properties, could theoretically be inhaled.
Camelids such as llamas naturally produce nanobodies when exposed to a virus, and Rout and colleagues have developed enormous libraries of promising SARS-CoV-2 nanobodies by giving a small dose of COVID protein to llamas (which produce nanobodies in response, much like humans produce antibodies in response to a vaccine). After taking small blood samples from the llamas and sequencing the nanobody DNA, the scientists later transfer key genes to bacteria, which in turn, produce many more nanobodies for lab analysis.
But screening these nanobody libraries to see how well they work (and which variants they work against) can be time-consuming and expensive. Rout and colleagues have long relied on the "mass spectrometry" technique, which works extraordinarily well but requires substantial expertise to perform and expensive equipment. They wondered whether a recently discovered "yeast display method", which was potentially far less expensive and simpler, could also effectively sort through their nanobody library.
Rout, in collaboration with Rockefeller's Fred Cross, started by first optimizing the yeast display method. (The two heads-of-lab took the unusual step of performing most of the benchwork themselves). They then used their optimized method to screen a library of nanobodies that they had previously screened with the mass spectrometry technique. They found that their version of the yeast display method not only identified many of the same nanobody candidates as the other approach, but also identified numerous other candidates that they had missed.
"The method is not ours," Cross clarifies. "But we made it simpler."
Toward nanobody therapy
The relatively simple and low-cost procedure described in the paper could empower laboratories in low-resource areas to generate nanobodies against SARS-CoV-2, as well as other viruses. "A researcher anywhere in the world, with fairly limited resources, could use this technique," Rout says. "The llama-related stuff could be FedEx-ed from North America."
For COVID, the long-term goal is that techniques such as these will lower the bar for entry into nanobody research and ultimately produce therapies that prevent infection. "How we'd make the therapeutic is unestablished, as yet," Cross says. "The specificity is there and the activity is there, but we don't have a drug yet. It'd be nice if we did. Hopefully someday."
Because with COVID now transitioning to an endemic disease, novel methods for preventing the infection cannot come soon enough. "New variants become prevalent by evading the immune system," Cross says. "It's important to have a fast way to find new nanobodies targeting the variants."
More information: Frederick R. Cross et al, Expanding and improving nanobody repertoires using a yeast display method: Targeting SARS-CoV-2, Journal of Biological Chemistry (2023). DOI: 10.1016/j.jbc.2023.102954
Journal information: Journal of Biological Chemistry
Provided by Rockefeller University