A better way to find RNA virus needles in database haystacks
A zoo once offered a coloring book featuring polar bears in winter scenes that came with crayons in various shades of white. To researchers searching for sequences of RNA viruses in large data sets, their work may be akin to finding a single snowflake on a colored-in page of that book.
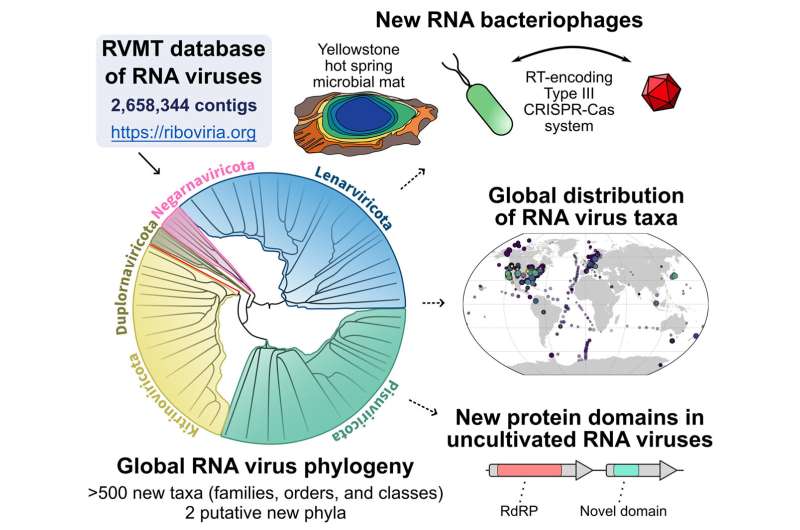
Published online September 28, 2022, in Cell, a team led by researchers at Tel Aviv University in Israel, the National Center for Biotechnology Information, and the U.S. Department of Energy (DOE) Joint Genome Institute (JGI), a DOE Office of Science User Facility located at Lawrence Berkeley National Laboratory (Berkeley Lab) describe a computational pipeline that can specifically scan for those snowflakes, or RNA virus sequences. Using this workflow, the team combed through more than 5,000 data sets of RNA sequences (metatranscriptomes) generated from diverse environmental samples around the world, resulting in a five-fold increase of RNA virus diversity.
"The world of viruses around us is vast, and we now have the means to explore it," said Eugene Koonin, a senior investigator at the NCBI and one of the senior authors on the paper, of the uncovered viral diversity. "Although the technical challenges of data analysis at this scale are formidable."
Computational sieves to filter sequences
There are more microbes on the planet than particles in a handful of dirt, and viruses vastly outnumber the microbes. Advances in sequencing technologies and computational tools have uncovered a diversity of viruses that infect not just crops, animals and humans, but also microbes whose presence or absence can impact the planet's nutrient cycles.
While most organism's genetic information is encoded in DNA, with RNA delivering the instructions inside DNA to the cell, RNA viruses store their genetic information in RNA without a DNA stage. "I would argue RNA viruses globally are even less known than DNA viruses," said Simon Roux, a JGI scientist and one of the project co-leads. "But same as DNA viruses, RNA viruses infect microbes all across the world and lead to cell death and/or profound changes in the cell physiology during infection."
While all RNA viruses have a gene that encodes for an enzyme called RNS-directed RNA Polymerase (RdRP), necessary for replicating the RNA genome replication, detecting it has been a challenge. Finding the RNA virus snowflakes in the snowstorm of genomic data involved developing special computational sieves to filter out sequences that were unlikely to contain the RdRP sequence.
The work resulted from a three-way collaboration that began in 2019, recalled Uri Neri of Tel Aviv University, one of the project co-leads and first author of the study. Members of the Tel Aviv and NCBI teams, who were already working on mining prokaryotic viruses together, learned from JGI's Nikos Kyrpides that his Microbiome Data Science group was also working on RNA virus mining. After a couple of virtual meetings of the three teams it was clear that a larger collaborative effort would be far more effective in achieving higher quality results compared to smaller individual efforts. This is also the type of synergistic and collaborative community spirit that the JGI advocates for and actively promotes.
The team used all the publicly available metatranscriptome datasets from the JGI's Integrated Microbial Genomes & Microbiomes (IMG/M) system. "We then looked into many more samples and refined our methodology," Neri said. "Our team grew and so did the scope of the project." To this end, Kyrpides emphasized, the contributions of the numerous JGI science users in collecting and submitting their microbiome samples for sequencing at the JGI cannot be overstated. Their cooperation and support, he said, and in several cases, their permission to use yet unpublished sequence data, was absolutely critical for the success of this effort and so was the acknowledgement of their contribution.
Both Roux and Koonin noted that the plethora of RNA virus sequences uncovered "significantly changes the global view of virus diversity," though not at the higher-level classifications of virus groups (phyla.) The new sequences are filling in some gaps on existing virus groups while also adding new branches. Additionally, RNA viruses do not appear to be evenly distributed around the world.
One expanded group is of viruses associated with bacteria; until now, most of the known RNA viruses have been associated with eukaryotes. Along with the expansion of bacteria-associated RNA viruses is the finding that "a few bacteria use CRISPR to defend against RNA," Roux noted, "although it's unclear why this is so rarely detected."
Developing approaches for reconciling 'real' Big Data
For the team, the computational work that led to the uncovered abundance of RNA viruses is just the beginning. "I often say that just identifying a sequence as viral is not even half the story." Neri said. "We invested a lot of our efforts into the post-discovery analyses—as best we could, we tried to describe the protein domains every virus carries, and who is their likely host. We've made all of that information fully free and openly available to the broader scientific community."
Uri Gophna from Tel Aviv University, and Koonin both noted that other research in parallel has reported similar "dramatic expansions" of the global RNA virome. "We now need to compare and reconcile the findings, coming up with a single, non-redundant dataset," said Koonin. "Hopefully, relatively soon we will be able to estimate the actual size of the RNA virome However, this is now real Big Data, we are dealing with billions of sequences, and soon, with trillions. The development of efficient, automated approaches to analyze and classify sequence data at this scale is essential."
More information: Uri Neri et al, Expansion of the global RNA virome reveals diverse clades of bacteriophages, Cell (2022). DOI: 10.1016/j.cell.2022.08.023
Journal information: Cell
Provided by DOE/Joint Genome Institute