Neural network speeds holographic image reconstruction for biological samples
Researchers have developed a new end-to-end neural network that can speed up the reconstruction of holographic images. Unlike other deep learning techniques, the approach can be used on samples not encountered during training, making it particularly useful for label-free holographic biomedical imaging.
"With this framework, a well-trained neural network can be distributed anywhere, without finetuning, and perform rapid and high-quality holographic imaging of various samples," explained research lead Hanlong Chen, University of California, Los Angeles (UCLA).
Hanlong Chen and Aydogan Ozcan will present the research at the Frontiers in Optics + Laser Science Conference (FiO LS) meeting being held in Rochester, New York and online 17—20 October 2022. The presentation is scheduled for Monday, 17 October at 16:30 EDT (UTC—04:00).
A generalizable approach
Although various neural networks have been developed to achieve the data-heavy task of hologram reconstruction for biological research and biomedical applications, most of them are designed to be very specific. This means that they may not perform well if used with samples that differ from those initially used to train the network.
To solve this problem, Chen and colleagues developed an end-to-end neural network called Fourier Imager Network (FIN). This type of neural network is trained using a single model, bypassing some of the steps usually used by other deep learning methods. End-to-end neural networks are also faster and potentially more generalizable to a wide variety of samples.
Faster, more accurate results
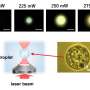
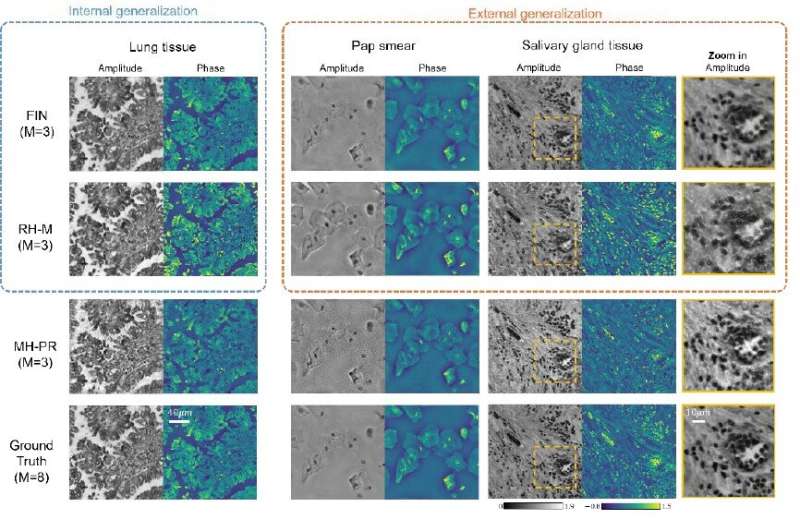
The FIN framework takes a sequence of intensity-only raw holograms captured at different sample-to-sensor distances with a lens-free in-line holographic microscope and creates reconstructed images of the samples. To test the new approach, the researchers trained the network using lung tissue sections. They then used FIN to reconstruct holographic images of human salivary gland tissue and Pap smear samples not seen by the network during training.
FIN worked well on these new types of samples and delivered more accurately reconstructed images than an iterative algorithm and a state-of-the-art deep learning model. It also showed an approximately 50-fold improved speed compared to the deep learning model. The researchers say that these results demonstrate the strong external generalization of FIN while also showing the immense potential of building broadly generalizable deep neural networks for various microscopy and computational imaging tasks.
Chen added, "Our next step is to investigate autofocusing while retaining the benefits of our approach, such as superb image quality, unprecedented generalization to new types of samples, and enhanced computational speed, making holographic imaging possible with low-resource devices."
Provided by Optica