Researchers unveil key processes in marine microbial evolution
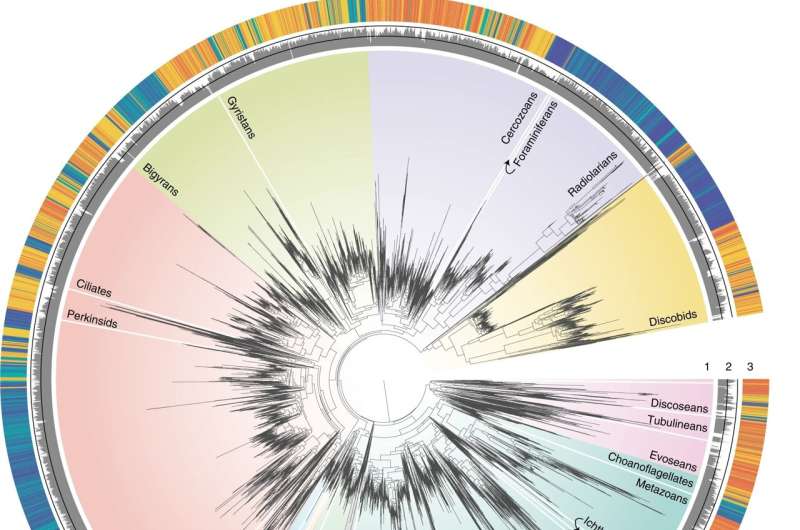
A study published recently in Nature Ecology and Evolution has unveiled some of the key processes in marine microbial evolution. According to the study, led by the Uppsala University (Sweden) and with the participation of the Institut de Ciències del Mar (ICM-CSIC) of Barcelona, it is the large number of habitat transitions -from sea to land and vice versa- that have occurred in the last millions of years that explains the great current diversity.
According to the authors, "crossing the salinity barrier is not easy for organisms and, when this happens, the resulting transitions are key evolutionary events that can trigger explosions of diversity." However, until now it was not known how frequent these transitions have been in the eukaryotic tree of life, which comprises animals, plants and a wide variety of eukaryotic microorganisms.
Small but very versatile
Specifically, the work published now has shown that microbial eukaryotes have made hundreds of great leaps from sea to land, and also to freshwater habitats, and vice versa, during their evolution. This, in turn, has made it possible to deduce where the ancestors of each of the microbial eukaryote groups were found.
"Thanks to the fact that we have good phylogenetic trees and samples from different environments, we have been able to analyze the habitat transitions in different groups of eukaryotes, which have been hundreds times during millions of years of eukaryotic evolution, which is more than we thought," explains Ramon Massana, ICM-CSIC researcher and one of the authors of the study.
For its elaboration, the scientific team used the latest technologies to sequence the DNA of microbes living in samples collected in boreal lakes, forest soils, the Indian Ocean and the Mariana Trench, among many other environments. Specifically, the ICM-CSIC provided marine samples obtained during the Malaspina expedition in different oceans and water column depths.
Thanks to this, it has been possible to construct large evolutionary trees of the organisms found in these environments and even to observe a series of patterns in the evolution of habitat preference.
"We found that organisms in the tree of life of eukaryotes are generally grouped according to whether they live in the oceans or in non-marine habitats," explains Mahwash Jamy, a researcher at Uppsala University and lead author of this study. In this regard, Jamy adds that "this finding confirms that adapting to a different salinity—or crossing the salt barrier—is difficult, even for microbes."
Nevertheless, the study proves that microbial eukaryotes have successfully established themselves in new habitats several hundred times throughout their evolution. Thus, the authors suggest that it is precisely these elusive transitions that would have allowed colonizing organisms to occupy vacant ecological niches, giving rise to the great diversity of eukaryotes today.
More clues about the first eukaryotes
On the other hand, evolutionary trees constructed from DNA sequences have also allowed researchers to zoom into the deep past and infer what the habitats of the ancestors of each microbial group might have been.
"It is likely that two of the largest groups of eukaryotes, the SARS and the Obozoa, each of which is larger than, for example, animals or plants, arose in completely different habitats," says Fabien Burki, also a researcher at Uppsala University and another of the study's lead authors.
According to Burki, the SAR lineage—which includes groups such as diatoms, ciliates, dinoflagellates, radiolarians, etc.—would have first arisen in the Precambrian oceans, while the ancestor of the Obazoa group -which has diversified into fungi, animals, choanoflagellates and amoebae- could have lived in non-marine habitats."
This shows, once again, that crossing the salinity barrier has played an important role in shaping eukaryotic evolution. For this reason, for future research, experts will turn to genomics to find out what genetic mechanisms underlie these key evolutionary events.
More information: Mahwash Jamy et al, Global patterns and rates of habitat transitions across the eukaryotic tree of life, Nature Ecology & Evolution (2022). DOI: 10.1038/s41559-022-01838-4
Journal information: Nature Ecology & Evolution
Provided by Institut de Ciències del Mar