Shining new light on cell membranes with improved imaging
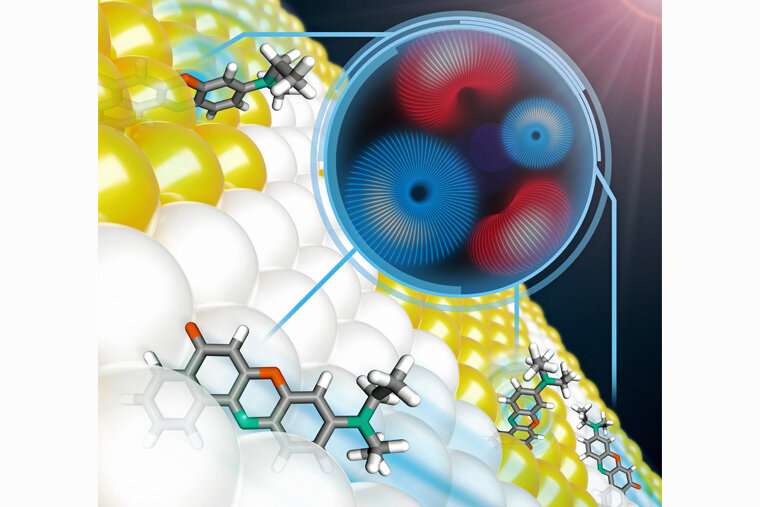
Research from the lab of Matthew Lew at Washington University in St. Louis offers entirely new ways to see the very small.
The research—two papers by Ph.D. students at the McKelvey School of Engineering—was published in the journals Optica and Nano Letters.
They have developed novel hardware and algorithms that allow them to visualize the building blocks of the biological world beyond three dimensions in a way that, until now, wasn't feasible. After all, cells are 3D objects and full of "stuff"—molecules—that moves around, rotates, spins and tumbles to drive life itself.
Like traditional microscopes, the work of two Ph.D. students in the Lew lab, Tingting Wu and Oumeng Zhang, uses light to peer into the microscopic world—but their innovations are anything but traditional. Currently, when people use light in imaging, they are likely interested in how bright that light is or what color it is. But light has other properties, including polarization.
"Oumeng's work twists the polarization of light," said Lew, assistant professor in the Preston M. Green Department of Electrical & Systems Engineering. "This way, you can see both how things translate (move in straight lines) and rotate at the same time"—something traditional imaging doesn't do.
"The development of new technology and the capability to see things we previously couldn't see is exciting," Zhang said. This unique capability to track both rotation and position at the same time gives him unique insights into how biological materials—human cells and pathogens, for instance—interact.
Wu's research also provides a new way to image cell membranes and, in a way, to see inside of them. Using fluorescent tracer molecules, she maps how the tracers interact with fat and cholesterol molecules in the membrane, determining how the lipids are arranged and organized.
"Any cell membrane, any nucleus, anything in the cell is a 3D structure," she said. "This helps us probe the full picture of a biological system. This enables us, for any biological sample, to see beyond three dimensions—we see the 3D structure plus three dimensions of molecular orientation, giving us 6D images."
The researchers developed computational imaging technology, which synergizes software and hardware together, to successfully see the previously unseeable.
"That's part of the innovation," Lew said. "Traditionally, biological imaging labs have been tied down to whatever commercial manufacturers are making. But if we engineer things differently, we can do so much more."
More information: Tingting Wu et al, Dipole-spread-function engineering for simultaneously measuring the 3D orientations and 3D positions of fluorescent molecules, Optica (2022). DOI: 10.1364/OPTICA.451899
Oumeng Zhang et al, Resolving the Three-Dimensional Rotational and Translational Dynamics of Single Molecules Using Radially and Azimuthally Polarized Fluorescence, Nano Letters (2022). DOI: 10.1021/acs.nanolett.1c03948
Journal information: Nano Letters , Optica
Provided by Washington University in St. Louis