Bonds from the past: A journey through the history of protein synthesis
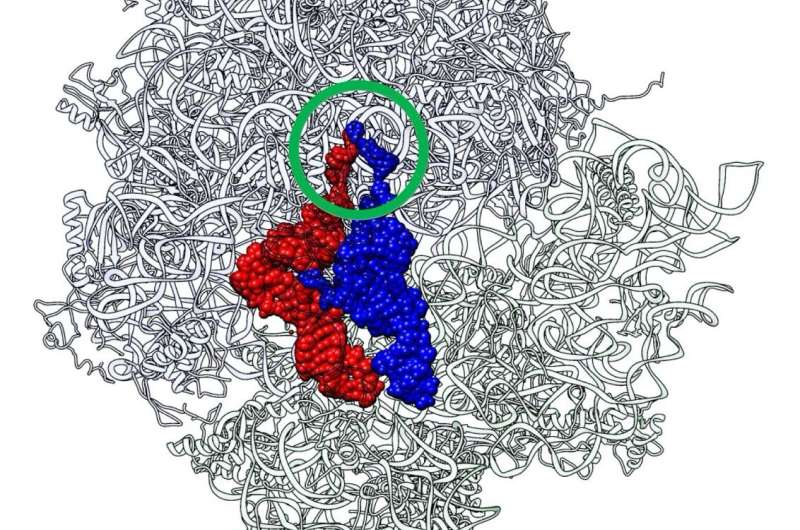
The genetic information stored in DNA is "decoded" to form proteins via the process of translation. This involves the formation of peptide bonds between amino acids bound to transfer RNA (tRNA) molecules that glide over the ribosome in very close proximity to each other, and elongate the peptide chain, which later undergoes conformational change, forming a protein. In contrast to the codon-dependent aminoacyl-tRNA recognition in the small ribosomal subunit, the peptide bond formation in question occurs at the peptidyl transferase center (PTC) of the large ribosomal subunit, in a non-amino acid specific manner. This non-specificity indicates that the large subunit evolved before the small subunit, which has more specific interactions with mRNA and tRNA.
Although the evolutionary process of PTC formation has been thoroughly documented, little is known about how ribosomes developed into functioning entities and became an essential component of protein synthesis. Scientists have long been perplexed by the fact that tRNAs require the help of a "scaffold" in order to create a peptide bond, which orients them for interaction via 3'-CCA sequences on their acceptor arms. What that scaffold is, and how it operates, would be intriguing to learn about.
A team of scientists at Tokyo University of Science, led by Prof. Koji Tamura, decided to solve this mystery using a perspective of continuity in biological evolution. Their study, which was published in the journal Life, sheds light on the evolutionary aspect of protein translation. Their results represent important evidence to prove the hypothesis about the origin and evolution of the PTC, which has changed the way we look at the modern-day ribosomes and tRNA.
The idea sprang to life after taking a close look at the crystal structure of the 70S ribosome-tRNA complex from Thermus thermophilus, a bacterium often used in the study of genetics. The peptidyl (P-) and aminoacyl (A-) sites of the tRNAs here aligned to bring the CCA termini in close proximity, like a rugby player's index fingers in the "Goromaru pose." "There was a certain entity that served as a scaffold for maintaining this proximity, and it most likely stemmed from the primordial PTC," says Prof. Tamura. Since an evolutionary aspect was likely, the team chose to utilize primordial tRNA or "RNA minihelix" for their study.
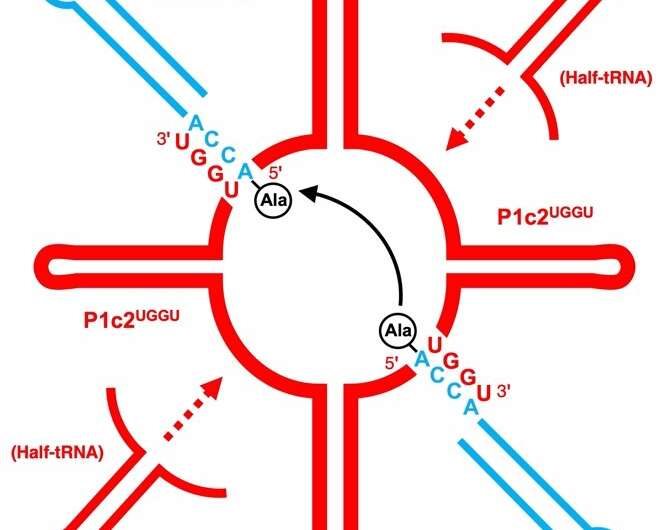
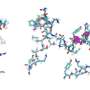
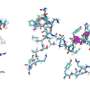
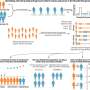
They first attempted a peptide bond formation between two alanine-specific minihelices in the presence of a ribosomal RNA segment. The peptide bond was formed using the ribosomal segment, P1c2, as an RNA scaffold which was just 70 nucleotides long! Next, they added a terminal amino acid segment (with the sequence UGGU) to the P1c2 (P1c2UGGU). According to mass spectrometry results, this increased the peptide bond formation ability by 4.2 times that of the original! The peptide bond formation between two alanine residues was supported by a scaffold of dimerized P1c2UGGU. The UGGU sequence of the scaffold interacted with the corresponding 3'-terminal ACCA of the minihelix and brought the two amino acids near enough to create a peptide bond. Nobel laureate Dr. Ada Yonath and her group recently showed that similar, conserved PTC regions could catalyze peptide bond formation with artificial analog molecules, but Prof. Tamura's group showed that an aminoacylated RNA could also be a substrate.
The findings definitely imply a possibility that minihelices bind to the primordial PTC. So, what do the results suggest about the evolution of ribosomes? "Functional interactions between the CCA of tRNA and PTC could have been 'revised' in the process of evolution. Although current ribosomes do not have a contiguous sequence like UGGU, their interactions are 'conceptually' similar to the effects seen in our study. It is plausible that minihelices eventually evolved into tRNA using, for example, kissing-loop interactions between two minihelix-like RNA molecules," Prof. Tamura explains. "These minihelix-like molecules, which form a part of the scaffold for peptide bond formation, may have not only contributed to the evolution of what is currently the PTC, but also formed tRNA molecules," he adds.
The future applications of this research—which has opened up exciting avenues in evolutionary RNA biology—are manifold. Faced with a metabolic paradox (that the components of DNA and RNA are generated from amino acids), it is conceivable to investigate the notion of "peptide nucleic acids" as genetic material precursors. The results are fascinating, and they will help scientists to decode molecular phenomena that have eluded them for years.
More information: Mai Kawabata et al, Peptide Bond Formation between Aminoacyl-Minihelices by a Scaffold Derived from the Peptidyl Transferase Center, Life (2022). DOI: 10.3390/life12040573
Provided by Tokyo University of Science