New study advances knowledge of the battle between viruses and human cells
In the long-term battle between a herpesvirus and its human host, a University of Massachusetts virologist and her team of students have identified some human RNA able to resist the viral takeover—and the mechanism by which that occurs.
This discovery, described in a paper published Feb. 15 in Proceedings of the National Academy of Sciences, represents an important step in the effort to develop anti-viral drugs to fight off infections.
"This paper is about trying to understand the mechanism that makes these RNA escape degradation," says senior author Mandy Muller, assistant professor of microbiology. "The next step is to figure out if we can manipulate this to our advantage."
In the Muller Lab, student researchers work with Muller studying how Kaposi sarcoma-associated herpesvirus (KSHV) hides for years inside the human body before seeking to gain control over human gene expression to complete the viral infection. At that point, people with a weakened immune system may develop Kaposi sarcoma cancer lesions in the mouth, skin or other organs.
The researchers use genome-wide sequencing, post-transcriptional sequencing and molecular biology to examine how the human cell or the virus knows how to prevent degradation.
"Viruses are very smart, that's what I love to say," Muller says. "They have lots of strategies to stick around, and they don't do a lot of damage for a very long time, because that's one way to hide from the immune system.
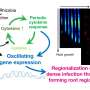
"But then, at some point—many, many years later—they reactivate. The way they do this is by triggering a massive RNA degradation event where the virus will wipe out the mRNA from the cell. That means the human system can no longer express the proteins that it needs to express, and that means also that a lot of resources are suddenly available for the virus."
How and why some RNA are able to escape the viral degradation are questions Muller's team—including lead author and graduate student Daniel Macveigh-Fierro and co-authors and undergraduates Angelina Cicerchia, Ashley Cadorette and Vasudha Sharma—has been investigating. The research was supported by a $1.9 million Maximizing Investigators' Research Award (MIRA) to Muller in 2020 from the NIH's National Institute for General Medical Sciences.
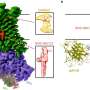
"We show that RNA that escape have a chemical tag on them—a post-transcriptional modification—that makes them different from the others," Muller explains. "By having this tag, M6A, they can recruit proteins that protect them from degradation."
Muller has been studying KSHV since she was an undergraduate in her native France, and her mission continues.
"We know you need this protein to protect the RNA from degradation, but we still don't know how that physically stops the degradation, so that's what we're going to look at now," she says.
Ultimately, understanding the mechanisms and pathways involved in KSHV infection may lead to the development of RNA therapeutics to treat viral diseases.
"By identifying the determinants of what makes an mRNA either resistant or susceptible to viral-induced decay, we could use those findings to our advantage to better design anti-viral drugs and reshape the outcome of infection," Muller says.
More information: Daniel Macveigh-Fierro et al, The m6A reader YTHDC2 is essential for escape from KSHV SOX-induced RNA decay, Proceedings of the National Academy of Sciences (2022). DOI: 10.1073/pnas.2116662119
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Massachusetts Amherst