Promoting the power of viral metagenomics
Since early 2020, people all over the world have been forced to confront the awesome power that viruses have to affect nearly every aspect of our daily lives. But for researchers who study viruses that infect organisms much smaller than humans, like bacteria, this power is not surprising at all. In fact, even in non-pandemic times such viruses are well known to play major roles in a vast range of globe-straddling phenomena. For example, viruses which infect the photosynthetic cyanobacteria found throughout the planet's oceans can exert great influence on earth's carbon and nitrogen cycles.
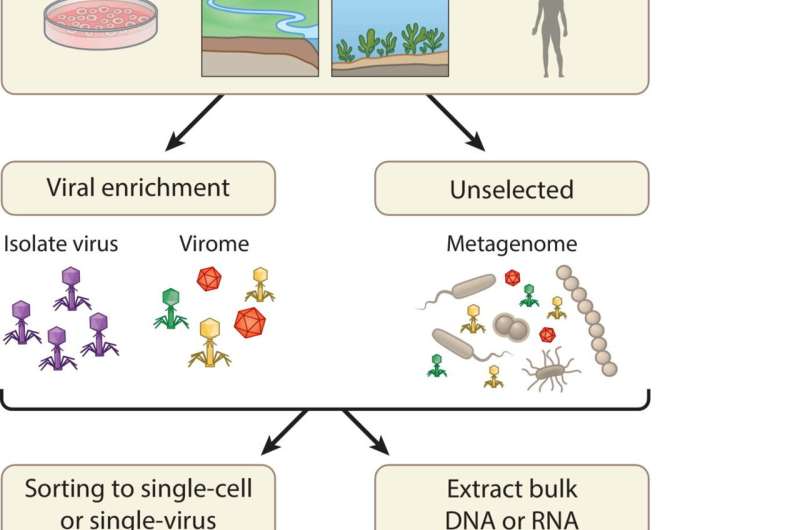
Currently one of the best ways to study such viruses is by sequencing the bulk DNA or RNA contained in a sample that has been collected from any kind of environment you can think of—animal intestines (including humans), grassland soils, even deep-sea hydrothermal vents—and computationally identifying which sequences represent viruses that were present in the sample. From these data it is possible to answer a wide variety of important questions such as which organisms the viruses may be targeting or infecting, how well the targeted organisms are at fending off the infection, and how long this evolutionary arms race has been going on. Answering these questions is critical to understanding the role of viruses in many areas such as ecology, agriculture, and medicine.
In the recently published article, the authors provide a brief history of the field, describe the most commonly used methods, and discuss some of the current challenges faced by those using metagenomics to try to understand the far-reaching impacts of viruses, which have been found in every biome on Earth.
More information: Lee Call et al, Illuminating the Virosphere Through Global Metagenomics, Annual Review of Biomedical Data Science (2021). DOI: 10.1146/annurev-biodatasci-012221-095114
Provided by DOE/Joint Genome Institute